Run parfor-Loops Without a Parallel Pool
This example shows how to run parfor-loops on a large cluster without a parallel pool.
Running parfor computations directly on a cluster allows you to use hundreds of workers to perform your parfor-loop. When you use this approach, parfor can use all the available workers in the cluster, and release the workers as soon as the loop completes. This approach is also useful if your cluster does not support parallel pools. However, when you run parfor computations directly on a cluster, you do not have access to DataQueue or Constant objects, and the workers restart between iterations, which can lead to significant overheads.
This example recreates the update of the ARGESIM benchmark CP2 Monte Carlo study [1] by Jammer et al [2]. For the CP2 Monte Carlo study, you simulate a spring-mass-damper system with different randomly sampled damping factors in parallel.
Create Cluster Object
Create the cluster object to and display the number of workers available in the cluster. HPCProfile is a profile for a MATLAB® Job Scheduler cluster.
cluster = parcluster("HPCProfile"); maxNumWorkers = cluster.NumWorkers; fprintf("Number of workers available: %d",maxNumWorkers)
Number of workers available: 496
Define Simulation Parameters
Set the simulation period, time interval, and initial states for the mass-spring system ODE.
period = [0 2]; % Use a period from 0 to 2 seconds h = 0.001; % time step t_interval = period(1):h:period(2); y0 = [0 0.1];
Set the number of iterations.
nReps = 10000000;
Initialize the random number generator and create an array of damping coefficients sampled from a uniform distribution with the range [800,1200].
rng(0); a = 800; b = 1200; d = (b-a).*rand(nReps,1)+a;
Run ODE Solver in Parallel
Initialize the results variable for the reduction operation.
y_sum = zeros(numel(t_interval),1);
Execute the ODE solver in a parfor-loop to simulate the system with varying damping coefficients. To run the parfor computations directly on the cluster, pass the cluster object as the second input argument to parfor. Use a reduction variable to compute the sum of the motion at each time step.
parfor(n = 1:nReps,cluster) f = @(t,y) massSpringODE(t,y,d(n)); [tOut,yOut] = ode45(f,t_interval,y0); y_sum = y_sum + yOut(:,1); end
Compute the mean response of the system and plot the response against time.
meanY = y_sum./numel(d); plot(t_interval,meanY) title("ODE Solution of Mass-Spring System") xlabel("Time") ylabel("Motion") grid on

Compare Computational Speedup
Compare the computational speedup of running the parfor-loop directly on the cluster to that of running the parfor-loop on a parallel pool.
Use the timeExecution helper function attached to this example to measure the execution time of the parfor-loop workflow on the client, on a parallel pool with 496 workers, and directly on a cluster with 496 workers available.
[serialTime,hpcPoolTime,hpcClusterTime] = timeExecution("HPCProfile",maxNumWorkers);
elapsedTimes = [serialTime hpcPoolTime hpcClusterTime];Calculate the computational speedup.
speedUp = elapsedTimes(1)./elapsedTimes;
fprintf("Speedup on cluster = %4.2f\nSpeedup on pool = %4.2f",speedUp(3),speedUp(2))Speedup on cluster = 154.11 Speedup on pool = 171.23
Create a bar chart comparing the speedup of each execution. The chart shows that running the parfor-loop directly on the cluster has a similar speedup to that of running the parfor-loop on a parallel pool.
figure; x = ["Client","Pool","Cluster"]; bar(x,speedUp); ylabel("Computational Speedup") xlabel("parfor Execution Environment") grid on
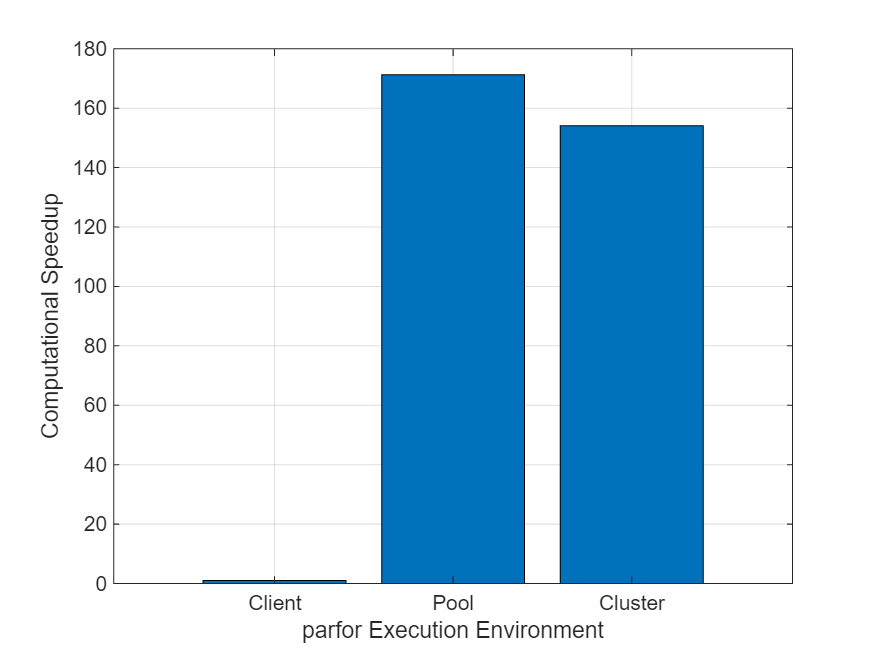
The speedup values are similar because the example uses a MATLAB Job Scheduler cluster. When you run the parfor-loop directly on a MATLAB Job Scheduler cluster, parfor can sometimes resuse workers without restarting them between iterations, which reduces overheads. If you run the parfor-loop directly on a third-party scheduler cluster, parfor restarts workers between iterations, which can result in significant overheads and much lower speedup values.
Helper Functions
This helper function represents the mass-spring system's ODEs that the solver uses.
You can rewrite the differential equation that describes the spring-mass system (eq1) as a system of first-order ODEs (eq2) that you can solve using the ode45 solver.
(eq1)
(eq2)
function dy = massSpringODE(t,y0,d) k = 9000; % spring stiffness (N/m) m = 450; % mass (kg) dy = zeros(2,1); dy(1) = y0(2); dy(2) = -(d*y0(2)+k*y0(1))/m; end
References
[1] Breitenecker, Felix, Gerhard Höfinger, Thorsten Pawletta, Sven Pawletta, and Rene Fink. "ARGESIM Benchmark on Parallel and Distributed Simulation." Simulation News Europe SNE 17, no. 1 (2007): 53-56.
[2] Jammer, David, Peter Junglas, and Sven Pawletta. “Solving ARGESIM Benchmark CP2 ’Parallel and Distributed Simulation’ with Open MPI/GSL and Matlab PCT - Monte Carlo and PDE Case Studies.” SNE Simulation Notes Europe 32, no. 4 (December 2022): 211–20. https://doi.org/10.11128/sne.32.bncp2.10625.
See Also
Scale Up Parallel Code to Large Clusters