particleoptions
Description
When you analyze a Bayesian nonlinear non-Gaussian state-space model (bnlssm model object),
particleoptions enables you to specify common sequential Monte Carlo
(SMC) sampler specifications through all stages of your workflow.
Creation
Description
options = particleoptionsbnlssm model object).
options = particleoptions(Property=Value)particleoptions(NumParticles=1e4,NewSamples="auxiliary") specifies
drawing 10,000 particles per sample time point from the proposal distribution and to use
the auxiliary SMC sampler.
Properties
Number of particles for SMC, specified as a positive integer.
Example: NumParticles=1e4
Data Types: double
Since R2025a
SMC proposal (importance) distribution, specified as a value in this table.
| Value | SMC Sampler | Description |
|---|---|---|
| Auxiliary filter [6] | SMC samples particles by using a reweight-resample strategy, which uses local approximations of the nonlinear state-space model and is informed by the observations. This option is
available only for models with observation noise (nonzero
|
"bootstrap" | Bootstrap forward filter [4] | SMC samples particles form the state transition distribution (observations do not inform the sampler). This option
is available only for models with observation noise (nonzero
This method has a relatively low computational cost, but it does not inform the routine of the current observations, which can increase the Monte Carlo sampling variance. |
"optimal" | Conditionally optimal proposal [2] | SMC samples particles from the one-step filtering distribution with incremental weights proportional to the likelihood (observations inform the sampler). The weights have minimum variance. This option is available for the following tractable models supplied in equation form:
|
"unscented" | Unscented transformation [7] | SMC proposal distribution is the one-step filtering distribution approximated by the unscented transformation (observations inform the sampler). This option is
available only for models in equation form with observation noise
(nonzero By default, this option
uses the filtered state mean as the representative point, around
which the algorithm generates sigma points. To specify all particles
as representative points for the unscented transformation, set
|
Example: NewSamples="unscented"
Data Types: char | string
SMC resampling method, specified as a value in this table.
| Value | Description |
|---|---|
"multinomial" | At time t, the set of previously generated particles (parent set) follows a standard multinomial distribution, with probabilities proportional to their weights. An offspring set is resampled with replacement from the parent set [1]. |
"residual" | Residual sampling, a modified version of multinomial resampling that can produce an estimator with lower variance than the multinomial resampling method [5]. |
"systematic" | Systematic sampling, which produces an estimator with lower variance than the multinomial resampling method [3]. |
Resampling methods downsample insignificant particles to achieve a smaller estimator variance than if no resampling is performed and to avoid sampling from a degenerate proposal [3].
Example: Resample="residual"
Data Types: char | string
Effective sample size threshold, below which particleoptions resamples particles, specified as a nonnegative scalar. For more details, see [3], Ch. 12.3.3.
Tip
To resample during every period, set
Cutoff=numparticles, wherenumparticlesis the value of theNumParticlesname-value argument.To avoid resampling, set
Cutoff=0.
Example: Cutoff=0.75*numparticles
Data Types: double
Examples
Create a default particle options object, and then adjust the default particle sampler and number of sampled particles. Then, pass it to a Bayesian nonlinear state-space model and fit the model to simulated data.
Consider this Bayesian nonlinear state-space model in equation.
where the parameters in have the following priors:
, that is, a truncated normal distribution with .
, that is, an inverse gamma distribution with shape and scale .
, that is, a gamma distribution with shape and scale .
Simulate Series
Consider this data-generating process (DGP).
where the series is a standard Gaussian series of random variables.
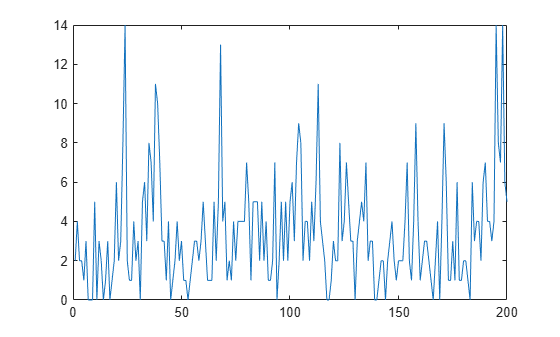
Simulate a series of 200 observations from the process.
rng(1,"twister") % For reproducibility T = 200; thetaDGP = [0.7; 0.2; 3]; numparams = numel(thetaDGP); MdlXSim = arima(AR=thetaDGP(1),Variance=thetaDGP(2), ... Constant=0); xsim = simulate(MdlXSim,T); y = random("poisson",thetaDGP(3)*exp(xsim)); figure plot(y)
Create and Adjust Default Particle Options Object
Create and display the default SMC particle sampler options by calling particleoptions without setting any inputs.
options = particleoptions; options
options =
particleoptions with properties:
NumParticles: 1000
NewSamples: "bootstrap"
Resample: "multinomial"
Cutoff: 500
options is a particleoptions object containing properties that specify particle sampling specifications for the SMC sampler.
Specify use of the auxiliary algorithm for sampling particles during SMC by setting the NewSamples property of options using dot notation.
options.NewSamples = "auxiliary";Create Bayesian Nonlinear Prior Model
A prior distribution is required to compose a posterior distribution. The Local Functions section contains the functions paramMap and priorDistribution required to specify the Bayesian nonlinear state-space model. The paramMap function specifies the state-space model structure and initial state moments. The priorDistribution function returns the log of the joint prior distribution of the state-space model parameters. You can use the functions only within this script.
Create a Bayesian nonlinear state-space model for the DGP.
Arbitrarily choose values for the hyperparameters.
Indicate that the state-space model observation equation is expressed as a distribution.
To speed up computations, the arguments
AandLogYof theparamMapfunction are written to enable simultaneous evaluation of the transition and observation densities of multiple particles. Specify this characteristic by using theMultipointname-value argument.Specify the SMC particle sampler options
options.
% pi(phi,sigma2) hyperparameters m0 = 0; v02 = 1; a0 = 1; b0 = 1; % pi(lambda) hyperparameters alpha0 = 3; beta0 = 1; hyperparams = [m0 v02 a0 b0 alpha0 beta0]; PriorMdl = bnlssm(@paramMap,@(x)priorDistribution(x,hyperparams), ... ObservationForm="distribution",Multipoint=["A" "LogY"],ParticleOptions=options);
PriorMdl is a bnlssm model specifying the state-space model structure and prior distribution of the state-space model parameters. Because PriorMdl contains unknown values, it serves as a template for posterior estimation with observations.
Choose Initial Parameter Values
Arbitrarily choose a set of initial parameter for the MCMC sampler.
theta0 = [0.5; 0.1; 2];
Estimate Posterior
Estimate the posterior by passing the prior model, simulated data, and initial parameter values to estimate.
PosteriorMdl = estimate(PriorMdl,y,theta0);
Optimization and Tuning
| Params0 Optimized ProposalStd
----------------------------------------
c(1) | 0.5000 0.6001 0.0979
c(2) | 0.1000 0.2166 0.0413
c(3) | 2 2.8669 0.2828
Posterior Distributions of Parameters
| Mean Std Quantile05 Quantile95
-----------------------------------------------
c(1) | 0.5951 0.0974 0.4193 0.7453
c(2) | 0.2522 0.0545 0.1690 0.3502
c(3) | 2.8667 0.2700 2.4043 3.2990
Posterior Distributions of Final States
| Mean Std Quantile05 Quantile95
-----------------------------------------------
x(1) | 0.4568 0.3541 -0.1100 1.0224
Proposal acceptance rate = 47.50%
PosteriorMdl.ParamMap
ans = function_handle with value:
@paramMap
ThetaPostDraws = PosteriorMdl.ParamDistribution; [numParams,numDraws] = size(ThetaPostDraws)
numParams = 3
numDraws = 1000
estimate finds an optimal proposal distribution for the MCMC sampler by using the tune function, and prints the optimal moments to the command line. estimate also displays a summary of the posterior distribution of the parameters. The true values of the parameters are close to their corresponding posterior means; all are within their corresponding 95% credible intervals.
PosteriorMdl is a bnlssm object representing the posterior distribution.
PosteriorMdl.ParamMapis the function handle to the function representing the state-space model structure; it is the same function asPrioirMdl.ParamMap.ThetaPostDrawsis a 3-by-1000 matrix of draws from the posterior distribution. By default,estimatetreats the first 100 draws as a burn-in sample and excludes them from the results.
To diagnose the MCMC sampler, create trace plots of the posterior parameter draws.
paramnames = ["\phi" "\sigma^2" "\lambda"]; figure h = tiledlayout(3,1); for j = 1:numParams nexttile plot(ThetaPostDraws(j,:)) hold on yline(thetaDGP(j)) ylabel(paramnames(j)) end title(h,"Posterior Trace Plots")
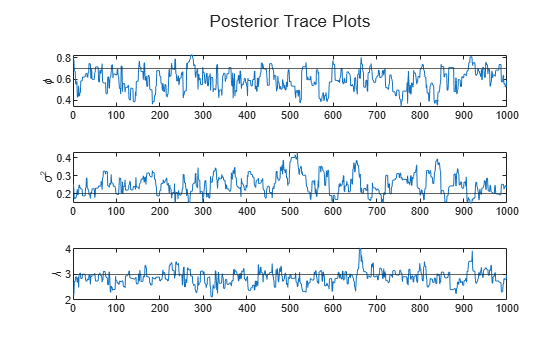
The sampler eventually settles at near the true values of the parameters. In this case, the sample shows serial correlation and transient behavior. You can remedy serial correlation in the sample by adjusting the Thin name-value argument, and you can remedy transient effects by increasing the burn-in period using the BurnIn name-value argument.
Local Functions
These functions specify the state-space model parameter mappings, in distribution form, and the log prior distribution of the parameters.
function [A,B,LogY,Mean0,Cov0,StateType] = paramMap(theta) A = theta(1); B = sqrt(theta(2)); LogY = @(y,x)y.*(x + log(theta(3)))- exp(x).*theta(3); Mean0 = 0; Cov0 = 2; StateType = 0; % Stationary state process end function logprior = priorDistribution(theta,hyperparams) % Prior of phi m0 = hyperparams(1); v20 = hyperparams(2); pphi = makedist("normal",mu=m0,sigma=sqrt(v20)); pphi = truncate(pphi,-1,1); lpphi = log(pdf(pphi,theta(1))); % Prior of sigma2 a0 = hyperparams(3); b0 = hyperparams(4); lpsigma2 = -a0*log(b0) - log(gamma(a0)) + (-a0-1)*log(theta(2)) - ... 1./(b0*theta(2)); % Prior of lambda alpha0 = hyperparams(5); beta0 = hyperparams(6); plambda = makedist("gamma",alpha0,beta0); lplambda = log(pdf(plambda,theta(3))); logprior = lpphi + lpsigma2 + lplambda; end
Specify particle sampler options for all SMC implementations of a Bayesian nonlinear state-space model.
Consider this Bayesian nonlinear state-space model and DGP in Default Particle Options Object.
Simulate a series of responses from the DGP.
rng(1,"twister") % For reproducibility T = 200; thetaDGP = [0.7; 0.2; 3]; numparams = numel(thetaDGP); MdlXSim = arima(AR=thetaDGP(1),Variance=thetaDGP(2), ... Constant=0); xsim = simulate(MdlXSim,T); y = random("poisson",thetaDGP(3)*exp(xsim));
Specify the following SMC particle sampler options:
The auxiliary algorithm for sampling particles
Sample 2,000 particles per time point
Residual particle resample method
options = particleoptions(NewSamples="auxiliary",Resample="residual");
Create a Bayesian nonlinear state-space prior model. Specify the SMC particle sampler options.
% pi(phi,sigma2) hyperparameters m0 = 0; v02 = 1; a0 = 1; b0 = 1; % pi(lambda) hyperparameters alpha0 = 3; beta0 = 1; hyperparams = [m0 v02 a0 b0 alpha0 beta0]; PriorMdl = bnlssm(@paramMap,@(x)priorDistribution(x,hyperparams), ... ObservationForm="distribution",Multipoint=["A" "LogY"],ParticleOptions=options);
Estimate the posterior by passing the prior model, simulated data, and initial parameter values to estimate.
theta0 = [0.5; 0.1; 2]; [PosteriorMdl,estParams] = estimate(PriorMdl,y,theta0);
Optimization and Tuning
| Params0 Optimized ProposalStd
----------------------------------------
c(1) | 0.5000 0.6989 0.0756
c(2) | 0.1000 0.1999 0.0544
c(3) | 2 2.9738 0.2521
Posterior Distributions of Parameters
| Mean Std Quantile05 Quantile95
-----------------------------------------------
c(1) | 0.5888 0.0970 0.4277 0.7494
c(2) | 0.2420 0.0489 0.1746 0.3325
c(3) | 2.8837 0.2812 2.3529 3.3596
Posterior Distributions of Final States
| Mean Std Quantile05 Quantile95
-----------------------------------------------
x(1) | 0.4796 0.3550 -0.1100 1.0591
Proposal acceptance rate = 35.40%
PosteriorMdl.ParticleOptions
ans =
particleoptions with properties:
NumParticles: 1000
NewSamples: "auxiliary"
Resample: "residual"
Cutoff: 500
estimate uses the particle sampler options in PriorMdl.ParticleOptions when it implements SMC.
Estimate smoothed state values from the posterior distribution.
states = smooth(PosteriorMdl,y,estParams,Method="marginal");smooth implements SMC to estimate states; it uses the options in PosteriorMdl.ParticleOptions to sample particles.
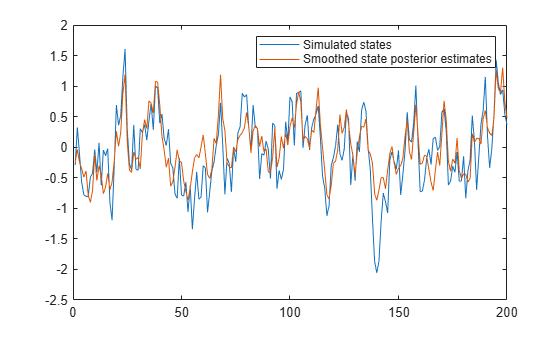
Visually compare the simulated states and smoothed state posterior estimates.
figure plot([xsim states]) legend(["Simulated states" "Smoothed state posterior estimates"])
Local Functions
These functions specify the state-space model parameter mappings, in distribution form, and the log prior distribution of the parameters.
function [A,B,LogY,Mean0,Cov0,StateType] = paramMap(theta) A = theta(1); B = sqrt(theta(2)); LogY = @(y,x)y.*(x + log(theta(3)))- exp(x).*theta(3); Mean0 = 0; Cov0 = 2; StateType = 0; % Stationary state process end function logprior = priorDistribution(theta,hyperparams) % Prior of phi m0 = hyperparams(1); v20 = hyperparams(2); pphi = makedist("normal",mu=m0,sigma=sqrt(v20)); pphi = truncate(pphi,-1,1); lpphi = log(pdf(pphi,theta(1))); % Prior of sigma2 a0 = hyperparams(3); b0 = hyperparams(4); lpsigma2 = -a0*log(b0) - log(gamma(a0)) + (-a0-1)*log(theta(2)) - ... 1./(b0*theta(2)); % Prior of lambda alpha0 = hyperparams(5); beta0 = hyperparams(6); plambda = makedist("gamma",alpha0,beta0); lplambda = log(pdf(plambda,theta(3))); logprior = lpphi + lpsigma2 + lplambda; end
References
[1] Andrieu, Christophe, Arnaud Doucet, and Roman Holenstein. "Particle Markov Chain Monte Carlo Methods." Journal of the Royal Statistical Society Series B: Statistical Methodology 72 (June 2010): 269–342. https://doi.org/10.1111/j.1467-9868.2009.00736.x.
[2] Doucet, Arnaud, Simon Godsill, and Christophe Andrieu. "On Sequential Monte Carlo Sampling Methods for Bayesian Filtering." Statistics and Computing 10 (July 2000): 197–208. https://doi.org/10.1023/A:1008935410038.
[3] Durbin, J, and Siem Jan Koopman. Time Series Analysis by State Space Methods. 2nd ed. Oxford: Oxford University Press, 2012.
[4] Gordon, Neil J., David J. Salmond, and Adrian F. M. Smith. "Novel Approach to Nonlinear/Non-Gaussian Bayesian State Estimation." IEE Proceedings F Radar and Signal Processing 140 (April 1993): 107–113. https://doi.org/10.1049/ip-f-2.1993.0015.
[5] Liu, Jun, and Rong Chen. "Sequential Monte Carlo Methods for Dynamic Systems." Journal of the American Statistical Association 93 (September 1998): 1032–1044. https://dx.doi.org/10.1080/01621459.1998.10473765.
[6] Pitt, Michael K., and Neil Shephard. "Filtering via Simulation: Auxiliary Particle Filters." Journal of the American Statistical Association 94 (June 1999): 590–599. https://doi.org/10.1080/01621459.1999.10474153.
[7] van der Merwe, Rudolph, Arnaud Doucet, Nando de Freitas, and Eric Wan. "The Unscented Particle Filter." Advances in Neural Information Processing Systems 13 (November 2000). https://dl.acm.org/doi/10.5555/3008751.3008833.
Version History
Introduced in R2026a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleziona un sito web
Seleziona un sito web per visualizzare contenuto tradotto dove disponibile e vedere eventi e offerte locali. In base alla tua area geografica, ti consigliamo di selezionare: .
Puoi anche selezionare un sito web dal seguente elenco:
Come ottenere le migliori prestazioni del sito
Per ottenere le migliori prestazioni del sito, seleziona il sito cinese (in cinese o in inglese). I siti MathWorks per gli altri paesi non sono ottimizzati per essere visitati dalla tua area geografica.
Americhe
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)