nlmefit
Fit nonlinear mixed-effects estimation
Syntax
Description
Examples
Load the sample data.
load orangetreedata.matThe vector c contains circumference measurements for five orange trees. The vector t contains the time at which the measurement was taken, and the vector id contains the tree id. The vector k contains the growth rate for each tree.
Create an anonymous function for the logistic growth equation.
model = @(phi,xfun,vfun)(phi(:,1))./(1+exp(-vfun(:).*(xfun-phi(:,2))));
model is a handle for a function given by the formula
.
The columns of the input argument phi correspond to the carrying capacity and midpoint for the logistic growth equation. xfun and vfun correspond to the time and logistic growth rate, respectively.
Fit model to the data in t, c, id, and k using the nlmefit function. Specify the initial estimate for each fixed-effect coefficient as 100.
beta = nlmefit(t,c,id,k,model,[100 100])
beta = 2×1
129.6589
121.4555
beta contains the fixed-effects estimates for the carrying capacity and midpoint.
Load the sample data.
load orangetreedata.matThe vector c contains circumference measurements for five orange trees. The vector t contains the time at which the measurement was taken, and the vector id contains the tree id. The vector k contains the growth rate for each tree.
Create an anonymous function for the logistic growth equation.
model = @(phi,xfun,vfun)(phi(:,1))./(1+exp(-vfun(:).*(xfun-phi(:,2))));
model is a handle for a function given by the formula
.
The columns of the input argument phi correspond to the carrying capacity and midpoint for the logistic growth equation. xfun and vfun correspond to the time and logistic growth rate, respectively.
Define an output function for nlmefit. For more information about the form of the output function, see the OutputFcn field description for the Options name-value argument.
function stop = outputFunction(beta,status,state) stop = 0; hold on plot3(status.iteration,beta(2),beta(1),"mo") state = string(state); if state=="done" stop=1; end end
outputFunction plots the iteration number for the fitting algorithm together with the fixed effects. outputFunction returns 1 when the fitting algorithm completes its final iteration.
Use the statset function to create an options structure for nlmefit that uses outputFunction as its output function.
default_opts=statset("nlmefit");
opts = statset(default_opts,OutputFcn=@outputFunction);opts is a statistics options structure that contains options for the fitting algorithm.
Create a figure and define axes in which to plot outputFunction. Fit model to the data in t, c, id, and k using the options in opts.
figure ax = axes(view=[12,10]); xlabel("Iteration") ylabel("beta(2)") zlabel("beta(1)") box on [beta,psi,stats] = nlmefit(t,c,id,k,model,[100 100],Options=opts)

beta = 2×1
129.6589
121.4555
psi = 2×2
154.9857 0
0 0.0000
stats = struct with fields:
dfe: 30
logl: -182.8139
mse: 1.9029e+03
rmse: 45.9892
errorparam: 43.6225
aic: 375.6279
bic: 373.6750
covb: [2×2 double]
sebeta: [9.7010 2.7699]
ires: [35×1 double]
pres: [35×1 double]
iwres: [35×1 double]
pwres: [35×1 double]
cwres: [35×1 double]
nlmefit calls outputFunction during the iterations of the fitting algorithm. The figure shows that the beta(1) and beta(2) fixed effects are near 126.5 and 129.6, respectively, when the iteration number is 0. The fitting algorithm calculates some estimates for beta(1) that are larger than 129.9 before converging to 129.66 in later iterations. Similarly, some estimates for beta(2) are larger than 122.2 in earlier iterations before converging to 121.5. The output argument beta contains the final values for the fixed effects. psi contains the covariance matrix for the random effects, and stats contains additional statistics about the fit.
Load the indomethacin data set.
load indomethacinThe variables time, concentration, and subject contain time series data for the blood concentration of the drug indomethacin in six patients.
Create an anonymous nonlinear function that accepts a vector of coefficients and a vector of predictor variables.
model = @(phi,t)(phi(1).*exp(-phi(2).*t)+phi(3).*exp(-phi(4).*t));
model is a handle for a function given by the formula
,
where is the concentration of indomethacin, for are coefficients, and is time. The function does not contain group-specific predictor variables because the formula does not include them.
Fit the model to the data using time as the predictor variable, subject as the grouping variable, and concentration as the response. Specify a log transformation function for the second and fourth coefficients.
phi0 = [1 1 1 1];
xform = [0 1 0 1];
[beta,psi,stats,b] = nlmefit(time,concentration, ...
subject,[],model,phi0,ParamTransform=xform)beta = 4×1
0.4606
-1.3459
2.8277
0.7729
psi = 4×4
0.0124 0 0 0
0 0.0000 0 0
0 0 0.3264 0
0 0 0 0.0250
stats = struct with fields:
dfe: 57
logl: 54.5882
mse: 0.0066
rmse: 0.0787
errorparam: 0.0815
aic: -91.1765
bic: -93.0506
covb: [4×4 double]
sebeta: [0.1092 0.2244 0.2558 0.1066]
ires: [66×1 double]
pres: [66×1 double]
iwres: [66×1 double]
pwres: [66×1 double]
cwres: [66×1 double]
b = 4×6
-0.1111 0.0871 0.0670 0.0115 -0.1315 0.0769
0 0 0 0 0 0
-0.7396 -0.0704 0.8004 -0.5654 0.4127 0.1624
0.0279 0.0287 0.0040 -0.2315 0.1984 -0.0276
The output argument beta contains the fixed effects for the model, and b contains the random effects. The maximum likelihood estimates for beta and the random-effects covariance matrix psi converge after about 300 iterations.
Plot the sample data together with the model, using only the fixed effects in beta for the model coefficients. Use the gscatter function to color code the data according to the subject. To reverse the log transformation on the second and fourth coefficients, take their exponentials using the exp function.
figure hold on gscatter(time,concentration,subject); phi = [beta(1),exp(beta(2)),beta(3),exp(beta(4))]; tt = linspace(0,8); cc = model(phi,tt); plot(tt,cc,LineWidth=2,Color="k") legend("Subject 1","Subject 2","Subject 3",... "Subject 4","Subject 5","Subject 6","Fitted curve") xlabel("Time (hours)") ylabel("Concentration (mcg/ml)") title("Indomethacin Elimination") hold off
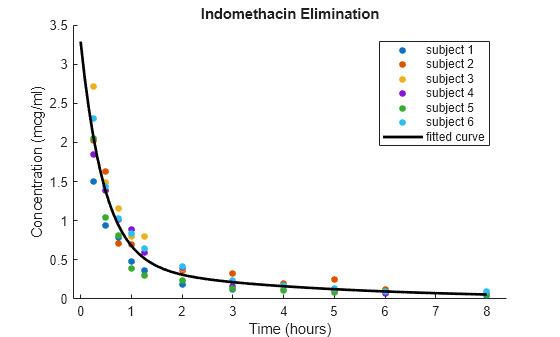
The plot shows that the blood concentration of indomethacin decreases over eight hours, and the fitted model passes through the bulk of the data.
Plot the data together with the model again, using both the fixed effects and the random effects in b for the model coefficients. For each subject, plot the data and the fitted curve in the same color.
figure hold on h = gscatter(time,concentration,subject); for j=1:6 phir = [beta(1)+b(1,j),exp(beta(2)+b(2,j)), ... beta(3)+b(3,j),exp(beta(4)+b(4,j))]; ccr = model(phir,tt); col = h(j).Color; plot(tt,ccr,Color=col) end legend("Subject 1","Subject 2","Subject 3",... "Subject 4","Subject 5","Subject 6",... "Fitted curve 1","Fitted curve 2","Fitted curve 3",... "Fitted curve 4","Fitted curve 5","Fitted curve 6") xlabel("Time (hours)") ylabel("Concentration (mcg/ml)") title("Indomethacin Elimination") hold off
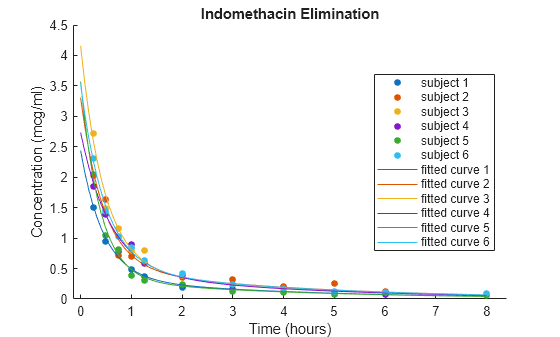
The plot shows that, for each subject, the fitted curve follows the bulk of the data more closely than the curve for the fixed-effects model in the previous figure. This result suggests that the random effects improve the fit of the model.
Input Arguments
Predictor variables, specified as an n-by-h numeric matrix, where n is the number of observations and h is the number of predictor variables.
Data Types: single | double
Response variable, specified as a numeric vector with n elements, where n is the number of observations.
Data Types: single | double
Grouping variable, specified as a numeric, categorical, or string vector, or a cell array of character vectors.
Example: ["subject1" "subject4" "subject3"…"subject3" "subject2"]
Example: [ones(50,1);2*ones(50,1);3*ones(50,1)]
Data Types: double | single | string | cell | categorical
Group predictor variables, specified as a numeric matrix or cell array. Group predictor variables have the same value for all observations in a group.
If the group-specific predictor variables are the same size across groups, specify
Vas an m-by-g matrix, where m is the number of groups and g is the number of group predictor variables.If the group-specific predictor variables vary in size across groups, specify
Vas an m-by-g cell array.If you do not have group predictor variables, specify
Vas[].
Example: [1 2; 2 4; 5 2; 7 1; 4 5; 3 6]
Data Types: single | double | cell
Model function, specified as a function handle. If
V is empty, the function can have the form yfit =
modelfun(phi,xfun). Otherwise, it must have the form yfit =
modelfun(phi,xfun,vfun).
| Argument | Description |
|---|---|
phi |
An array of model coefficients. The size of
|
xfun |
An array of predictor variables. The size of
|
vfun |
An array of group predictor variables. The size of
If |
yfit | Fitted response values. yfit contains an element for each row in
xfun. |
For improved performance, allow modelfun to accept predictor
variables for multiple observations and specify the Vectorization name-value
argument as "SingleGroup" or "Full".
Example: model =
@(phi,xfun,vfun)(phi(:,1).*exp(vfun(:,1).*xfun(:,1))+log(phi(:,2).*xfun(:,2)./vfun(:,1)));
beta = nlmefit(X,y,group,V,model,[1 1 1]);
Data Types: function_handle
Initial fixed-effects values, specified as a numeric vector or matrix.
If
beta0is a vector, its size must be equal to the number of fixed effects.If
beta0is a matrix, it must have as many rows as the number of model coefficients. For each column inbeta0,nlmefitrepeats the estimation of the fixed effects using the column entries as the initial fixed-effects values.
Example: [1 1 1 1]
Data Types: single | double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: nlmefit(X,y,group,v,model,[1 1
1],Vectorization="SingleGroup") specifies for nlmefit
to estimate the model function parameters twice using the same starting point, and to pass
the sample data to the model function in batches corresponding to each group.
Fixed and Random Effects
Fixed-effects coefficients, specified as a numeric or logical vector. This argument indicates which coefficients in the model function input argument phi contain a fixed effect. For more information about the model function, see modelfun. By default, all the elements of phi contain fixed effects.
When you specify FEParamsSelect, you cannot specify
FEConstDesign, FEGroupDesign, or
FEObsDesign.
Example: FEParamsSelect=[1 0 1 1]
Data Types: single | double | logical
Fixed-effects
design matrix, specified as a p-by-f numeric matrix.
p is the number of coefficients in the model function input argument
phi, and f is the number of fixed effects. For more
information, see modelfun and Mixed-Effects Model Hierarchy.
When you specify FEConstDesign, you cannot specify
FEParamsSelect, FEGroupDesign, or
FEObsDesign.
Example: FEConstDesign=[0.1 0.5; 3 0.7; 0.8 2; 4 8] specifies a
design matrix for four model parameters and two fixed effects.
Data Types: single | double
Fixed-effects group design matrices, specified as a
p-by-f-by-m array of numeric
matrices. p is the number of coefficients in the model function input
argument phi, f is the number of fixed effects, and
m is the number of groups specified by group.
FEGroupDesign specifies a different
p-by-f design matrix for each of the
m groups.
When you specify FEGroupDesign, you cannot specify
FEConstDesign, FEParamsSelect, or
FEObsDesign.
Example: A(:,:,1) = [0.1 0.5; 3 0.7; 0.8 2]; A(:,:,2) = ones(3,2);
FEGroupDesign=A specifies a design matrix for three model parameters, two
fixed effects, and two groups.
Data Types: single | double
Fixed-effects observation design matrices, specified as a
p-by-f-by-n array of
numeric matrices. p is the number of coefficients in the model
function input argument phi, f is the number of
fixed effects, and n is the number of observations specified by
X and y. FEObsDesign
specifies a different p-by-f design matrix for
each of the n observations.
When you specify FEObsDesign, you cannot specify
FEGroupDesign, FEConstDesign, or
FEParamsSelect.
Example: A(:,:,1) = [0.1 0.5; 3 0.7; 0.8 2];...; A(:,:,n) = ones(3,2);
FEObsDesign=A specifies a design matrix for three model parameters, two
fixed effects, and n observations.
Data Types: single | double
Random-effects coefficients, specified as a numeric or
logical vector. This argument indicates which coefficients in the model function input argument
phi contain a random effect. For more information about the model
function, see modelfun. By default, all the elements of
phi contain random effects.
When you specify REParamsSelect, you cannot specify
REConstDesign, REGroupDesign, or
REObsDesign.
Example: REParamsSelect=[0 0 1 1]
Data Types: single | double | logical
Random-effects design matrix, specified as a p-by-r
numeric matrix. p is the number of coefficients in the model function input
argument phi, and r is the number of random effects. For
more information, see modelfun and Mixed-Effects Model Hierarchy.
When you specify REConstDesign, you cannot specify
REParamsSelect, REGroupDesign, or
REObsDesign.
Example: REConstDesign=[1 0 2 1; 3 1 0 0; 0 0 1 0] specifies a
design matrix for three parameters and four random effects.
Data Types: single | double
Random-effects group design matrices, specified as a
p-by-r-by-m array of
numeric matrices. p is the number of coefficients in the model
function input argument phi, r is the number of
random effects, and m is the number of groups specified by
group. REGroupDesign specifies a different
p-by-r design matrix for each of the
m groups.
When you specify REGroupDesign, you cannot specify
REConstDesign, REParamsSelect, or
REObsDesign.
Example: A(:,:,1) = [0.1 0.5; 3 0.7; 0.8 2]; A(:,:,2) = ones(3,2);
REGroupDesign=A specifies a design matrix for three model parameters, two
random effects, and two groups.
Data Types: single | double
Random-effects observation design matrices, specified as a
p-by-r-by-n array of
numeric matrices. p is the number of coefficients in the model
function input argument phi, r is the number of
random effects, and n is the number of observations specified by
X and y. REObsDesign
specifies a different p-by-r design matrix for
each of the n observations.
When you specify REObsDesign, you cannot specify
REGroupDesign, REConstDesign, or
REParamsSelect.
Example: A(:,:,1) = [0.1 0.5; 3 0.7; 0.8 2];...; A(:,:,n) = ones(3,2);
REObsDesign=A specifies a design matrix for three model parameters, two
fixed effects, and n observations.
Data Types: single | double
Iterative Algorithm
Method for approximating the loglikelihood of the model, specified as one of the following:
"LME"— Loglikelihood for the linear mixed-effects model at the current conditional estimates ofbetaandb"RELME"— Restricted loglikelihood for the linear mixed-effects model at the current conditional estimates ofbetaandb"FO"— First-order Laplacian approximation without random effects"FOCE"— First-order Laplacian approximation at the conditional estimates ofb
The function calculates the conditional estimates for
beta and b using the current estimates of
the parameters in the covariance matrix for b.
Example: ApproximationType="FO"
Data Types: string | char
Parameterization for the scaled covariance matrix, specified as one of the following:
"logm"— Matrix logarithm"chol"— Cholesky factorization
Example: CovParameterization="chol"
Data Types: string | char
Random-effects covariance matrix pattern, specified as an
r-by-r identity, numeric, or logical matrix,
or a 1-by-r vector of integers, where r is the
number of random effects. nlmefit calculates variance and
covariance estimates for the random effects specified by
CovPattern. For more information, see
psi.
When
CovPatternis a matrix, each off-diagonal element corresponds to a pair of random effects, and each element along the diagonal corresponds to the variance for a single random effect.nlmefitcalculates variance and covariance estimates for the random effects corresponding to the nonzero elements ofCovPattern. Zero elements inCovPatternindicate that the variances and covariances are constrained to zero.If
CovPatternis not a row-column permutation of a block diagonal matrix,nlmefitautomatically adds nonzero elements to produce such a pattern.When
CovPatternis a vector, elements with equal values define groups of random effects. In addition to variances,nlmefitcalculates covariance estimates for pairs of random effects within groups, and constrains covariances across groups to zero.
The default value for CovPattern is an
r-by-r identity matrix, which corresponds to
uncorrelated random effects.
Example: CovPattern=ones(3)
Example: CovPattern=[1 1 2]
Data Types: single | double | logical
Model for the error term, specified as a string scalar or character vector containing the
model name. nlmefit stores the parameter values for the error
model in the errorparam field of the stats
output argument.
| Model Name | Formula | stats.errorparam |
|---|---|---|
"constant" |
| a |
"proportional" |
| b |
"combined" |
| [a,b] |
"exponential" |
| a |
In the above table, is the response variable y, is the fitted model, and and are model parameters.
Example: ErrorModel="exponential"
Data Types: string | char
Optimization function for maximizing the likelihood function, specified as
"fminsearch" or "fminunc".
"fminsearch"— Uses a direct search method involving function evaluations only. For more information, seefminsearch."fminunc"— Uses gradient methods. This option, which requires Optimization Toolbox, is generally more efficient. For more information, seefminunc(Optimization Toolbox).
For more information about the nlmefit fitting algorithm,
see Algorithms.
Example: OptimFun="fminunc"
Data Types: string | char
Options, specified as a statistics options structure returned by the statset function. The structure contains the following fields.
| Field | Description |
|---|---|
DerivStep | Relative difference used in the finite difference gradient
calculation, specified as a positive scalar, or as a vector of the same
length as the |
Display | Option to display algorithm information, specified as one of the following:
|
FunValCheck | Flag to check for invalid values returned by the model function, specified as one of the following:
|
OutputFcn | Output function, specified as a function handle or a cell array
of function handles.
The default output function plots the progress of the fitting algorithm for estimating the fixed effects, and plots the random-effects variances. |
Streams | Streams used by nlmefit when generating random
numbers, specified as a RandStream object. The default is
the current global stream. |
TolX | Termination tolerance for the fixed- and random-effects estimates. The
default is 1e-4. |
TolFun | Termination tolerance for the objective function value, specified
as a positive scalar. This setting is only available for censored data.
The default is |
MaxIter | Maximum number of iterations allowed, specified as a positive
integer. The default is |
For more information about the nlmefit fitting algorithm,
see Algorithms.
Example: Options=statset("nlmefit")
Data Types: struct
Coefficient transformation function for the model function, specified as a vector of
integers between 0 and 3, inclusive. ParamTransform must have the same
number of elements as the model function input argument phi. For more
information about the model function, see modelfun.
The coefficients for the model function are given by
where and are the original and transformed model coefficients, respectively. and are the design matrices for the fixed and random effects and , respectively. The value of ParamTransform indicates
the form of the transformation function according to the following scheme.
| Value | Function |
|---|---|
0 |
|
1 |
|
2 |
|
3 |
|
For more information about the fixed-effects design matrix, see
FEConstDesign and FEGroupDesign. For more
information about the random-effects design matrix, see
REConstDesign and REGroupDesign.
Example: ParamTransform=[0 1 0]
Data Types: single | double
Flag to refine the initial fixed-effects values, specified as
"on" or "off". When
RefineBeta0 is "on",
nlmefit first fits the model without random effects, and
then replaces beta0 with the fitted fixed effects.
Example: RefineBeta0="off"
Data Types: string | char
Size of the model function inputs, specified as a string scalar or character vector
with one of the following values. For more information about the model function inputs,
see modelfun.
| Value | Description |
|---|---|
"SinglePhi" |
The
model function calculates |
"SingleGroup" |
The model function calculates
|
"Full" |
The model function calculates
|
If V is an empty array,
nlmefit calls the model function with two inputs.
Example: Vectorization="SingleGroup"
Data Types: string | char
Output Arguments
Fixed-effects coefficient estimates, returned as a numeric vector.
Data Types: single | double
Random-effects covariance matrices, returned as an r-by-r numeric matrix.
Data Types: single | double
Fitting information, returned as a structure with the following fields.
| Field | Description |
|---|---|
logl | Maximized loglikelihood for the fitted model. |
rmse | Square root of the estimated error variance.
nlmefit calculates rmse on the
log scale when the ErrorModel name-value argument is
"exponential" during fitting. |
errorparam | Estimated parameters of the error model specified by the
ErrorModel name-value argument during fitting. |
aic | Akaike information criterion (AIC). nlmefit
calculates the AIC as aic = -2*logl+2*numParam, where logl is the maximized
loglikelihood for the fitted model, and numParam is the
number of fitting parameters. The fitting parameters include the degrees of
freedom for the random-effects covariance matrix, the number of fixed effects,
and the number of parameters in the error model. |
bic | Bayesian information criterion (BIC).
|
sebeta | Standard errors for the fixed-effects estimates in
beta. |
covb | Estimated covariance matrix for the fixed-effects estimates in
|
dfe | Error degrees of freedom |
pres | Population residuals, given by |
ires | Individual residuals, given by |
pwres | Population weighted residuals |
cwres | Conditional weighted residuals |
iwres | Individual weighted residuals |
Data Types: struct
Random-effects estimates, returned as an
r-by-m numeric matrix. r is
the number of random effects and m is the number of groups specified
by group.
Data Types: single | double
Algorithms
nlmefit fits the model by maximizing an estimate of the marginal
likelihood with the random effects removed by integration. During fitting, the function makes
the following assumptions:
The random effects come from a multivariate normal distribution and are independent between groups.
The observation errors come from the same normal distribution, and are independent of the random effects and each other. To change this default assumption, specify the
ErrorModelname-value argument.
References
[1] Lindstrom, M. J., and D. M. Bates. “Nonlinear Mixed-Effects Models for Repeated Measures Data.” Biometrics. Vol. 46, 1990, pp. 673–687.
[2] Davidian, M., and D. M. Giltinan. Nonlinear Models for Repeated Measurements Data. New York: Chapman & Hall, 1995.
[3] Pinheiro, J. C., and D. M. Bates. “Approximations to the Log-Likelihood Function in the Nonlinear Mixed-Effects MSodel.” Journal of Computational and Graphical Statistics. Vol. 4, 1995, pp. 12–35.
[4] Demidenko, E. Mixed Models: Theory and Applications. Hoboken, NJ: John Wiley & Sons, Inc., 2004.
Version History
Introduced in R2008b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleziona un sito web
Seleziona un sito web per visualizzare contenuto tradotto dove disponibile e vedere eventi e offerte locali. In base alla tua area geografica, ti consigliamo di selezionare: .
Puoi anche selezionare un sito web dal seguente elenco:
Come ottenere le migliori prestazioni del sito
Per ottenere le migliori prestazioni del sito, seleziona il sito cinese (in cinese o in inglese). I siti MathWorks per gli altri paesi non sono ottimizzati per essere visitati dalla tua area geografica.
Americhe
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)