Al momento, stai seguendo questo contributo
- Vedrai gli aggiornamenti nel tuo feed del contenuto seguito
- Potresti ricevere delle email a seconda delle tue preferenze per le comunicazioni
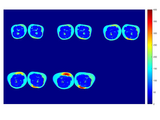
These scripts allow you to create parametric mapping of 3D MRI data and extract the parameters:T1,T2,perfusion constant, fast apparent diffusion coefficient, diffusion constant and apparent diffusion coefficient.
You will need to download these as a supplement:
nifti libraries: https://uk.mathworks.com/matlabcentral/fileexchange/8797-tools-for-nifti-and-analyze-image
threshold: https://uk.mathworks.com/matlabcentral/fileexchange/29372-thresholding-an-image
file sort: https://uk.mathworks.com/matlabcentral/fileexchange/47434-natural-order-filename-sort
Cita come
matt birkbeck (2026). Parametric Mapping Scripts for MRI data (https://it.mathworks.com/matlabcentral/fileexchange/64579-parametric-mapping-scripts-for-mri-data), MATLAB Central File Exchange. Recuperato .
Riconoscimenti
Ispirato da: Tools for NIfTI and ANALYZE image, Thresholding an image, Natural-Order Filename Sort
Informazioni generali
- Versione 1.2.0.0 (236 KB)
Compatibilità della release di MATLAB
- Compatibile con qualsiasi release
Compatibilità della piattaforma
- Windows
- macOS
- Linux