clustergram
Object containing hierarchical clustering analysis data
Description
The clustergram function creates a
clustergram object. The object contains hierarchical clustering analysis
data that you can view in a heatmap and dendrogram.
Creation
Description
cgObj =
clustergram( performs hierarchical
clustering analysis on the values in data)data. The returned clustergram
object cgObj contains analysis data and displays a dendrogram and
heatmap.
cgObj =
clustergram(
sets the object properties using
name-value pairs. For example, data,Name,Value)clustergram(data,'Standardize','column')
standardizes the values along the columns of data. You can specify multiple name-value
pairs. Enclose each property name in quotes.
Input Arguments
Source data, specified as a DataMatrix object or numeric matrix. Typically, if the matrix contains gene expression data, each row corresponds to a gene and each column corresponds to a sample.
Name-Value Arguments
Use comma-separated name-value pair arguments to set the object properties. Enclose each property name in single quotes.
Example: cg =
clustergram(data,'Colormap',redbluecmap,'Annotate',true)
Properties
Dimension for standardizing data values, specified as a character vector, string, or positive integer. Choices are:
'column'or1— Standardize along the columns of data.'row'or2— Standardize along the rows of data.'none'or3— Do not standardize.
If you specify 'column' or 'row', the function transforms the standardized values so that the mean is 0 and the standard deviation is 1 in the specified dimension.
Example: 'column'
Data Types: double | char | string
Flag to make the heatmap color scale symmetric around zero, specified as
true or false.
Example: false
Data Types: logical
Name of a function or function handle to impute missing data, specified as a character vector or cell array. If you specify a cell array, the first element must be the name of a function or function handle, and the remaining elements must be name-value pairs used as inputs to the function. Missing data points are colored gray in the heatmap.
If data points are missing, use this property to impute the missing values..
Otherwise, the clustergram function errors.
Example: 'func1'
Data Types: char
heatmap colors, specified as a three-column (M-by-3) matrix of
red-green-blue (RGB) values or the name of a function handle that returns a colormap,
such as redgreencmap or redbluecmap.
The default colormap is redgreencmap, in which red represents
values above the mean, black represents the mean, and green represents values below the
mean of a row (gene) across all columns (samples).
Example: redbluecmap
Data Types: double | char
Column labels, specified as a string vector, cell array of character vectors, or
numeric vector. The size of the vector must match the number of columns in the input
data.
If the number of column labels is 200 or more, the labels do not appear in the clustergram plot.
Example: ["sample1","sample2","sample3"]
Data Types: double | string | cell
Row labels, specified as a string vector, cell array of character vectors, or
numeric vector. The size of the vector must match the number of rows in the input
data.
If the number of row labels is 200 or more, the labels do not appear in the clustergram plot.
Example: ["gene1","gene2","gene3"]
Data Types: double | string | cell
Orientation of column labels, specified as a numeric scalar. Specify the value of rotation in degrees (positive angles cause counterclockwise rotation).
Example: 30
Data Types: double
Orientation of row labels, specified as a numeric scalar. Specify the value of rotation in degrees (positive angles cause counterclockwise rotation).
Example: 30
Data Types: double
Flag to display data values in the heatmap, specified as true or false.
Example: true
Data Types: logical
Display precision of data values in the heatmap, specified as a numeric scalar. The default number of digits of precision is 2.
Example: 3
Data Types: double
Flag to display colored markers instead of colored text for the row and column labels,
specified as true or false.
Example: true
Data Types: logical
Text color of displayed data values in the heatmap, specified as a character vector,
string, or three-element numeric vector. For example, to use cyan, you can enter
[0 1 1], 'c', "c",
"cyan", or 'cyan'. For details, see Color Options.
Example: 'red'
Data Types: char | string | double
Display range of standardize values, specified as a positive scalar.
The default value 3means that there is a color variation for
values between -3 and 3, but values greater than
3 are the same color as 3, and values less than
-3 are the same color as -3.
For example, if you specify redgreencmap for the
'Colormap' property, pure red represents values greater than or
equal to the specified display range value and pure green represents values less than or
equal to the negative of the specified display range value.
Example:
3
Data Types: double
Warning
This property will be removed in a future release. Set
LabelsWithMarkers to true for colored
markers instead of colored texts.
Color information for column labels, specified as a structure or structure array.
For a single structure, you must specify the following fields.
Labels— Cell array of character vectors specifying column labels listed in theColumnLabelsproperty.Colors— Character vector or string specifying a color for the column labels. If this field is empty, the default color (black) is used.
For a structure array, you must specify a single element in each field for each structure.
Labels— Character vector or string specifying a column label listed in theColumnLabelsproperty.Colors— Character vector or string specifying a color for the column labels. If this field is empty, the default color (black) is used.
For more information on specifying colors, see Color Options.
Data Types: struct
Warning
This property will be removed in a future release. Set
LabelsWithMarkers to true for colored
markers instead of colored texts.
Color information for row labels, specified as a structure or structure array.
For a single structure, you must specify the following fields.
Labels— Cell array of character vectors specifying row labels listed in theRowLabelsproperty.Colors— Character vector or string specifying a color for the row labels. If this field is empty, the default color (black) is used.
For a structure array, you must specify a single element in each field for each structure.
Labels— Character vector or string specifying a row label listed in theRowLabelsproperty.Colors— Character vector or string specifying a color for the row labels. If this field is empty, the default color (black) is used.
For more information on specifying colors, see Color Options.
Dimension for data clustering, specified as a positive integer, character vector, or string. Choices are:
'column'or1— Cluster along the columns of data only, which results in clustered rows.'row'or2— Cluster along the rows of data only, which results in clustered columns.'all'or3— Cluster along the columns of data, then cluster along the rows of row-clustered data.
Example: 2
Data Types: double | char | string
Information for annotating groups of columns, specified as a structure or structure array.
If you specify a single structure, each field must contain a cell array of elements. If you specify a structure array, each structure must have a single element in each field.
The fields are :
GroupNumber— Scalar specifying the column group number to annotate.Annotation— Character vector specifying text to annotate the column group.Color— Character vector or three-element vector of RGB values specifying a color to label the column group. For more information on specifying colors, see Color Options. If this field is empty, the default value is'blue'.
Data Types: struct
Distance metric to pass to the pdist function to calculate the pairwise distances between columns,
specified as a character vector or cell array. Specify a cell array if the distance
metric requires extra arguments. For example, to use the Minkowski distance with an
exponent p, specify
{'minkowski',p}.
Example: 'jaccard'
Data Types: char | cell
Color threshold information to pass to the dendrogram function to create a dendrogram plot, specified as a scalar,
two-element numeric vector, character vector, or cell array of character vectors. This
option sets the 'ColorThreshold' property of the dendrogram plot. If
you specify a two-element numeric vector or cell array, the first element is for the
rows, and the second element is for the columns.
Data Types: double | cell
Ratio of space that the row and column dendrograms occupy relative to the heatmap,
specified as a scalar between 0 and 1 or
two-element vector. If you specify a scalar, the function uses it as the ratio for both
row and column dendrograms. If you specify a two-element vector, the function uses the
first element for the ratio of the row dendrogram width to the heatmap width, and the
second element for the ratio of the column dendrogram height to the heatmap height. The
second element is ignored for one-dimensional clustergrams.
Example: 0.5
Data Types: double
Linkage method passed to the linkage function to create the hierarchical cluster tree for rows and
columns, specified as a character vector or two-element cell array of character vectors.
If you specify a cell array, the function uses the first element for linkage between
rows, and the second element for linkage between columns.
Example: 'centroid'
Data Types: char | cell
Flag to log2 transform the data from natural scale, specified
as true or false.
Example: true
Data Types: logical
Flag to calculate the optimal leaf order that maximizes the similarity between
neighboring leaves, specified as true or false.
The default value depends on the size of the input data. If the
number of rows or columns in data exceeds 1500, the default value
is false. Otherwise, the default value is
true.
Disabling the optimal leaf ordering calculation can be useful when working with large datasets because this calculation consumes a lot of memory and time.
Example: true
Data Types: logical
Information for annotating groups of rows, specified as a structure or structure array.
If you specify a single structure, each field must contain a cell array of elements. If you specify a structure array, each structure must have a single element in each field.
The fields are
GroupNumber— Scalar specifying the column group number to annotate.Annotation— Character vector specifying text to annotate the column group.Color— Character vector or three-element vector of RGB values specifying a color to label the column group. For more information on specifying colors, see Color Options. If this field is empty, the default value is'blue'.
Data Types: struct
Distance metric to pass to the pdist function to calculate the pairwise distances between rows, specified
as a character vector or cell array. Specify a cell array if the distance metric
requires extra arguments. For example, to use the Minkowski distance with an exponent
p, specify
{'minkowski',p}.
Example: 'jaccard'
Data Types: char | cell
Flag to show the dendrogram tree diagrams with the clustergram, specified as
'on' or 'off'.
Example: 'off'
Data Types: char
Object Functions
Examples
Load microarray data containing gene expression levels of Saccharomyces cerevisiae (yeast) during the metabolic shift from fermentation to respiration [1].
load filteredyeastdata
This MAT file includes three variables, which are added to the MATLAB® workspace:
- yeastvalues - A matrix of gene expression data from Saccharomyces -_cerevisiae_ during the metabolic shift from fermentation to respiration - genes - A cell array of GenBank® accession numbers for labeling the rows in yeastvalues - times - A vector of time values for labeling the columns in yeastvalues
Create a clustergram object to display the heat map from the gene expression data in the first 30 rows of the yeastvalues matrix and standardize along the rows of data.
cgo = clustergram(yeastvalues(1:30,:),'Standardize','Row')
Clustergram object with 30 rows of nodes and 7 columns of nodes.

Use the set method and the genes and times vectors to add meaningful row and column labels to the clustergram.
set(cgo,'RowLabels',genes(1:30),'ColumnLabels',times)

Add a color bar to the clustergram by clicking the Insert Colorbar button on the toolbar.
View a data tip containing the intensity value, row label, and column label for a specific area of the heat map by clicking the Data Cursor button on the toolbar, then clicking an area in the heat map. To delete this data tip, right-click it, then select Delete Current Datatip.
Display intensity values for each area of the heat map by clicking the Annotate button on the toolbar. Click the Annotate button again to remove the intensity values.
Tip: If the amount of data is large enough, the cells within the clustergram are too small to display the intensity annotations. Zoom in to see the intensity annotations.
Remove the dendrogram tree diagrams from the figure by clicking the Show Dendrogram button on the toolbar. Click it again to display the dendrograms.
Use the get method to display the properties of the clustergram object, cgo.
get(cgo)
Cluster: 'ALL'
RowPDist: {'Euclidean'}
ColumnPDist: {'Euclidean'}
Linkage: {'Average'}
Dendrogram: {}
OptimalLeafOrder: 1
LogTrans: 0
DisplayRatio: [0.2000 0.2000]
RowGroupMarker: []
ColumnGroupMarker: []
ShowDendrogram: 'on'
Standardize: 'ROW'
Symmetric: 1
DisplayRange: 3
Colormap: [11×3 double]
ImputeFun: []
ColumnLabels: {1×7 cell}
RowLabels: {30×1 cell}
ColumnLabelsRotate: 90
RowLabelsRotate: 0
Annotate: 'off'
AnnotPrecision: 2
AnnotColor: 'w'
ColumnLabelsColor: []
RowLabelsColor: []
LabelsWithMarkers: 0
Change the clustering parameters by changing the linkage method and changing the color of the groups of nodes in the dendrogram whose linkage is less than a threshold of 3.
set(cgo,'Linkage','complete','Dendrogram',3)
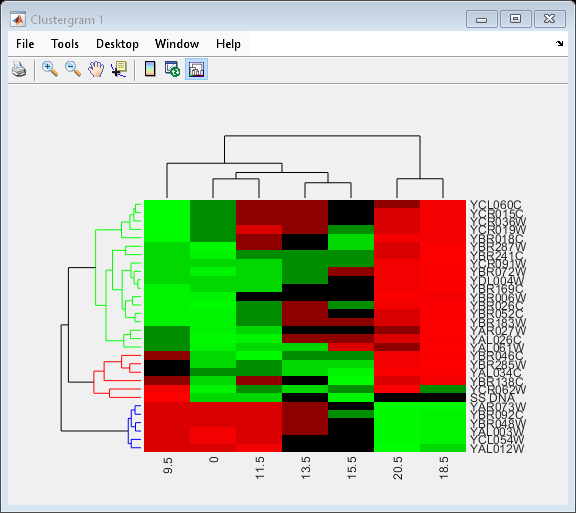
Place the cursor on a branch node in the dendrogram to highlight (in blue) the group associated with it. Press and hold the mouse button to display a data tip listing the group number and the nodes (genes or samples) in the group.

Right-click a branch node in the dendrogram to display a menu of options.

The following options are available:
- Set Group Color - Change the cluster group color. - Print Group to Figure - Print the group to a figure window. - Copy Group to New Clustergram - Copy the group to a new clustergram window. - Export Group to Workspace - Create a clustergram object of the group in the MATLAB workspace. - Export Group Info to Workspace - Create a structure containing information about the group in the MATLAB workspace. The structure contains these fields:
- GroupNames - Cell array of character vectors containing the names of the row or column groups. - RowNodeNames - Cell array of character vectors containing the names of the row nodes. - ColumnNodeNames - Cell array of character vectors containing the names of the column nodes. - ExprValues - An M-by-N matrix of intensity values, where M and N are the number of row nodes and of column nodes respectively. If the matrix contains gene expression data, typically each row corresponds to a gene and each column corresponds to sample.
Create a clustergram object for Group 18 in the MATLAB workspace. Right-click Group 18, then select Export Group to Workspace. In the Export to Workspace dialog box, type Group18, then click OK.
Use the view method to view the clustergram object, Group18.
view(Group18)

View all the gene expression data using a diverging red and blue colormap and standardize along the rows of data.
cgo_all = clustergram(yeastvalues,'Colormap',redbluecmap,'Standardize','Row')
Clustergram object with 614 rows of nodes and 7 columns of nodes.

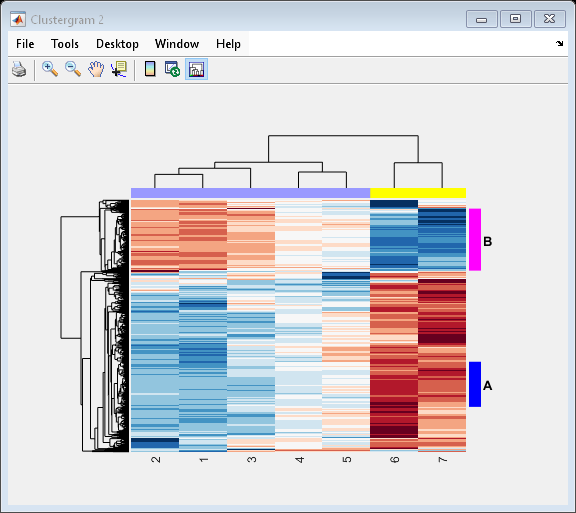
Create structure arrays to specify marker colors and annotations for two groups of rows (510 and 593) and two groups of columns (4 and 5).
rm = struct('GroupNumber',{510,593},'Annotation',{'A','B'},... 'Color',{'b','m'}); cm = struct('GroupNumber',{4,5},'Annotation',{'Time1','Time2'},... 'Color',{[1 1 0],[0.6 0.6 1]});
Use the RowGroupMarker and ColumnGroupMarker properties to add the color markers and annotations to the clustergram.
set(cgo_all,'RowGroupMarker',rm,'ColumnGroupMarker',cm)
More About
The following lists the predefined colors and their RGB triplet equivalents. The short names and long names are character vectors that specify one of eight preset colors. The RGB triplet is a three-element row vector whose elements specify the intensities of the red, green, and blue components of the color; the intensities must be in the range [0 1].
RGB Triplet | Short Name | Long Name |
|---|---|---|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
References
[1] DeRisi, J. L. “Exploring the Metabolic and Genetic Control of Gene Expression on a Genomic Scale.” Science 278, no. 5338 (October 24, 1997): 680–86.
Version History
Introduced before R2006a
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleziona un sito web
Seleziona un sito web per visualizzare contenuto tradotto dove disponibile e vedere eventi e offerte locali. In base alla tua area geografica, ti consigliamo di selezionare: .
Puoi anche selezionare un sito web dal seguente elenco:
Come ottenere le migliori prestazioni del sito
Per ottenere le migliori prestazioni del sito, seleziona il sito cinese (in cinese o in inglese). I siti MathWorks per gli altri paesi non sono ottimizzati per essere visitati dalla tua area geografica.
Americhe
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)