bioformatsread
Description
Add-On Required: This feature requires the Medical Imaging Toolbox Interface for Whole Slide Imaging File Reader add-on.
bim = bioformatsread(filename)filename using the Bio-Formats
library and returns the main image data series as a blockedImage
object or array of blockedImage objects.
bim = bioformatsread(filename,Name=Value)ImageType="thumbnail" reads the associated thumbnail image,
if present in the file.
Examples
Run this code to download a whole slide image from the MathWorks® website and unzip the downloaded folder. The image, stored in the NDPI format, contains an H&E stained microscopy slide provided by the OpenSlide library test data set [1].
zipFile = matlab.internal.examples.downloadSupportFile("image","data/CMU-1.zip"); filepath = fileparts(zipFile); if ~exist(filepath) unzip(zipFile,filepath) end filename = fullfile(filepath,"CMU-1.ndpi");
Read the image data by using the Bio-Formats library. The function returns a blockedImage object.
im = bioformatsread(filename);
The blocked image contains multiple resolution levels, which is common for whole slide images, as it helps to manage performance and avoid out-of-memory issues. View the number of resolution levels by checking the NumLevels property.
im.NumLevels
ans = 4
Display the image by using the imageshow function. The function picks the resolution level to display based on the current viewer size and your screen size. As you zoom in, the resolution level automatically adjusts to a finer resolution.
imageshow(im)
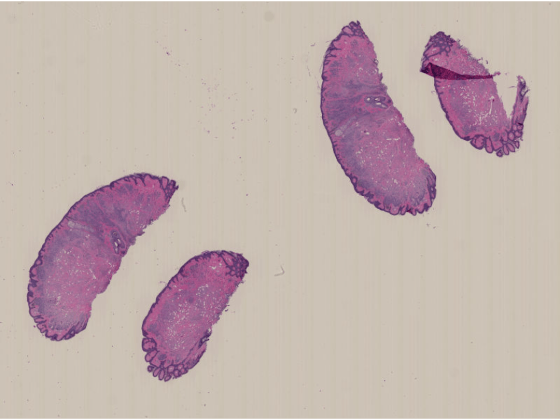
Run this code to download a whole slide image (WSI) from the MathWorks website and unzip the downloaded folder. The image, stored in the NDPI format, contains an H&E stained microscopy slide provided by the OpenSlide library test data set [1].
zipFile = matlab.internal.examples.downloadSupportFile("image","data/CMU-1.zip"); filepath = fileparts(zipFile); if ~exist(filepath) unzip(zipFile,filepath) end filename = fullfile(filepath,"CMU-1.ndpi");
Read the metadata from the file. The function returns a structure, info, that contains fields for the filename, vendor name, associated images, and raw metadata.
info = bioformatsinfo(filename);
Check the AssociatedImages field to see if the file includes a label, macro, or thumbnail image. The file used in this example includes a macro image.
info.AssociatedImages
ans = "macro image"
Read the macro image by specifying the ImageType argument. If you specify an image type that is not present in the file, the function returns an error.
macroIm = bioformatsread(filename,ImageType="macro");Display the macro image. The macro image provides an overview of the slide, including the label.
imageshow(macroIm)
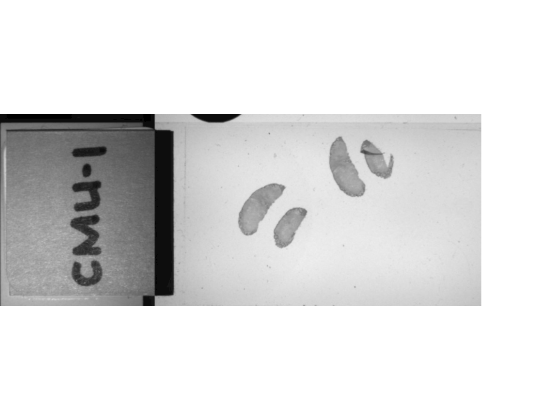
Run this code to download a whole slide image from the MathWorks website and unzip the downloaded folder. The image is available in the OpenSlide library test data set [2]. The file is in the CZI format, which is associated with ZEISS microscope cameras. The image shows trichrome stain on a mouse kidney sample.
zipFile = matlab.internal.examples.downloadSupportFile("image","data/Zeiss-5-JXR.zip/Zeiss-5-JXR.zip"); zipPath = fileparts(zipFile); filepath = fileparts(zipPath); if ~exist(filepath) unzip(zipFile,filepath) end filename = fullfile(filepath,"Zeiss-5-JXR.czi");
Read the file metadata. The function returns a structure, info, that contains metadata about the file.
info = bioformatsinfo(filename);
View the SeriesImages value, which indicates the file contains two image series corresponding to two different scan regions.
info.SeriesImages
ans = 2×1 string array
"scanregion0"
"scanregion1"
The core metadata indicates that all images in the file are 2-D RGB images with one time point. There are 12 rows, indicating a total of 12 resolution levels across the image series and associated images.
info.RawMetadata.CoreInfo
ans=1×12 struct array with fields:
11323 11338 1 3 1 "uint8" 1 "XYCZT"
5661 5669 1 3 1 "uint8" 1 "XYCZT"
2830 2834 1 3 1 "uint8" 1 "XYCZT"
1415 1417 1 3 1 "uint8" 1 "XYCZT"
707 708 1 3 1 "uint8" 1 "XYCZT"
11300 11335 1 3 1 "uint8" 1 "XYCZT"
5650 5667 1 3 1 "uint8" 1 "XYCZT"
2825 2833 1 3 1 "uint8" 1 "XYCZT"
1412 1416 1 3 1 "uint8" 1 "XYCZT"
706 708 1 3 1 "uint8" 1 "XYCZT"
640 515 1 3 1 "uint8" 1 "XYCZT"
1260 615 1 3 1 "uint16" 1 "XYCZT"
Read the image series from the file. By default, the bioformatsread function reads all image series, excluding any associated images. If a file contains multiple image series, the function returns them as an array of blockedImage objects, where each object represents one series.
bimSeries = bioformatsread(filename)
bimSeries =
1×2 blockedImage array with properties:
Read only properties
Source: 'Various'
Adapter: [1×1 wsi.blocked.BioFormatsAdapter]
ClassUnderlying: [5×1 string]
Settable properties
No properties.
View the number of resolution levels for the first series.
bimSeries(1).NumLevels
ans = 5
Display the first series by using the imageshow function.
imageshow(bimSeries(1))
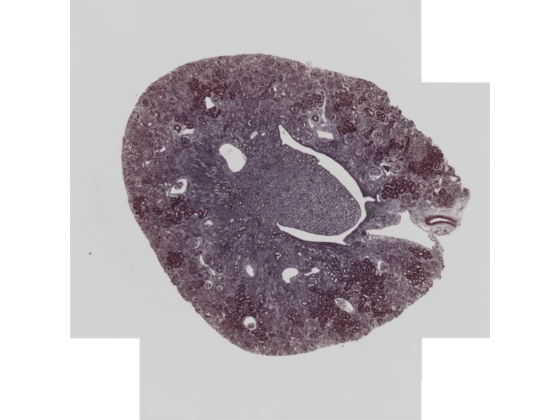
Specify the name of an OME-TIFF file containing a simulated multi-time point whole slide image (WSI) provided by The Open Microscopy Environment [3]. The image is attached to this example as a supporting file.
filename = fullfile(pwd,"multi-channel-time-series.ome.tif");Read the metadata from the file.
info = bioformatsinfo(filename);
View the RawMetadata.CoreInfo field, which shows that the image contains one 2-D image with three channels and seven time points.
info.RawMetadata.CoreInfo
ans = struct with fields:
SizeX: 439
SizeY: 167
SizeZ: 1
SizeC: 3
SizeT: 7
PixelType: "int8"
ImageCount: 21
DimensionOrder: "XYZCT"
By default, the bioformatsread function reads the image from the first time point in a series.
imIdx1 = bioformatsread(filename);
To read a different time point, specify the TimeSeriesIdx argument. Read the third time point.
imIdx3 = bioformatsread(filename,TimeSeriesIdx=3);
Input Arguments
Name of the file, specified as a string scalar or a character vector. Specify
filename as the absolute path to the file, a relative path from
the current directory, or a relative path from a directory on the MATLAB® path.
The file must be in a format supported by the Bio-Formats library. For a list of supported formats, see the Supported Formats page in the Bio-Formats documentation.
Data Types: char | string
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: bioformatsread(filename,ImageType="thumbnail") reads the
associated thumbnail image, if present in the file.
Type of image to read, specified as one of the values in the table.
| Value | Description |
|---|---|
"data" | Read the main image data. You can access the number
and names of the main image data series in the
|
"label" | Read the associated label image. If the file does
not contain a label image, the function returns an error. You can check
whether the file contains a label image using the
|
"macro" | Read the associated macro image. If the file does not
contain a macro image, the function returns an error. You can check
whether the file contains a macro image using the
|
"thumbnail" | Read the associated thumbnail image. If the file does not contain a thumbnail image, the function returns an error. You can
check whether the file contains a thumbnail image using the
|
Data Types: char | string
Time series index to read, specified as a positive integer. Specify a value that
corresponds to a valid time point in your image data. For example, if your image file
contains three time points, valid values include 1,
2, and 3.
If specified in the file metadata, the number of time points is accessible in the
RawMetadata.CoreInfo.SizeT field of the structure returned by
bioformatsinfo.
Example: TimeSeriesIdx=2
Data Types: single | double | int8 | int16 | int32 | int64 | uint8 | uint16 | uint32 | uint64
Output Arguments
Image data, returned as a blockedImage
object or as an array of blockedImage objects. The function returns a
scalar blocked image when the main data image or the associated image specified by
ImageType contains one series. The function returns an array of
blocked images when reading multiseries image data.
References
[1] "CMU‑1.ndpi". OpenSlide Test Data. Accessed November 10, 2025. https://openslide.cs.cmu.edu/download/openslide-testdata/Hamamatsu/.
[2] Venklab, Pathology, UT Health San Antonio. "Zeiss‑5‑JXR.czi". OpenSlide Test Data. Accessed November 10, 2025. https://openslide.cs.cmu.edu/download/openslide-testdata/Zeiss/.
[3] "multi-channel-time-series.ome.tif". Open Microscopy Environment Sample Images. Accessed November 10, 2025. https://downloads.openmicroscopy.org/images/OME-TIFF/2016-06/bioformats-artificial/.
Version History
Introduced in R2026a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleziona un sito web
Seleziona un sito web per visualizzare contenuto tradotto dove disponibile e vedere eventi e offerte locali. In base alla tua area geografica, ti consigliamo di selezionare: .
Puoi anche selezionare un sito web dal seguente elenco:
Come ottenere le migliori prestazioni del sito
Per ottenere le migliori prestazioni del sito, seleziona il sito cinese (in cinese o in inglese). I siti MathWorks per gli altri paesi non sono ottimizzati per essere visitati dalla tua area geografica.
Americhe
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)