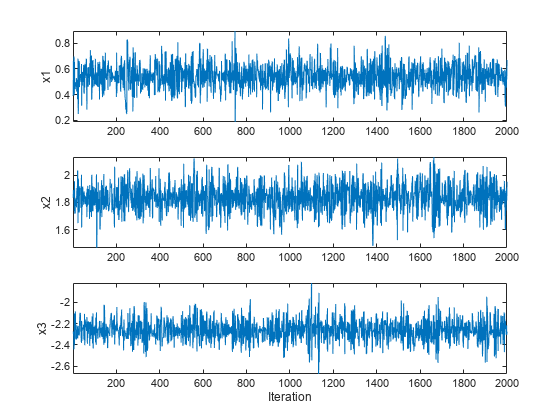
drawSamples
Class: HamiltonianSampler
Generate Markov chain using Hamiltonian Monte Carlo (HMC)
Syntax
chain = drawSamples(smp)
[chain,endpoint,accratio]
= drawSamples(smp)
[chain,endpoint,accratio]
= drawSamples(___,Name,Value)
Description
chain = drawSamples(smp)smp.
[ also returns the
final state of the Markov chain in chain,endpoint,accratio]
= drawSamples(smp)endpoint and
the fraction of accepted proposals in accratio.
[ specifies
additional options using one or more name-value pair arguments. Specify
name-value pair arguments after all other input arguments.chain,endpoint,accratio]
= drawSamples(___,Name,Value)
Input Arguments
Name-Value Arguments
Output Arguments
Examples
Tips
After creating an HMC sampler using the
hmcSamplerfunction, you can compute MAP (maximum-a-posteriori) point estimates, tune the sampler, draw samples, and check convergence diagnostics using the methods of theHamiltonianSamplerclass. For an example of this workflow, see Bayesian Linear Regression Using Hamiltonian Monte Carlo.
Version History
Introduced in R2017a