fit
Train linear model for incremental learning
Description
The fit function fits a configured incremental learning model for linear regression (incrementalRegressionLinear object) or linear binary classification (incrementalClassificationLinear object) to streaming data. To additionally track performance metrics using the data as it arrives, use updateMetricsAndFit instead.
To fit or cross-validate a regression or classification model to an entire batch of data at once, see the other machine learning models in Regression or Classification.
Examples
Create a default incremental linear SVM model for binary classification. Specify an estimation period of 5000 observations and the SGD solver.
Mdl = incrementalClassificationLinear('EstimationPeriod',5000,'Solver','sgd')
Mdl =
incrementalClassificationLinear
IsWarm: 0
Metrics: [1×2 table]
ClassNames: [1×0 double]
ScoreTransform: 'none'
Beta: [0×1 double]
Bias: 0
Learner: 'svm'
Properties, Methods
Mdl is an incrementalClassificationLinear model. All its properties are read-only.
Mdl must be fit to data before you can use it to perform any other operations.
Load the human activity data set. Randomly shuffle the data.
load humanactivity n = numel(actid); rng(1) % For reproducibility idx = randsample(n,n); X = feat(idx,:); Y = actid(idx);
For details on the data set, enter Description at the command line.
Responses can be one of five classes: Sitting, Standing, Walking, Running, or Dancing. Dichotomize the response by identifying whether the subject is moving (actid > 2).
Y = Y > 2;
Fit the incremental model to the training data, in chunks of 50 observations at a time, by using the fit function. At each iteration:
Simulate a data stream by processing 50 observations.
Overwrite the previous incremental model with a new one fitted to the incoming observations.
Store , the number of training observations, and the prior probability of whether the subject moved (
Y=true) to see how they evolve during incremental training.
% Preallocation numObsPerChunk = 50; nchunk = floor(n/numObsPerChunk); beta1 = zeros(nchunk,1); numtrainobs = zeros(nchunk,1); priormoved = zeros(nchunk,1); % Incremental fitting for j = 1:nchunk ibegin = min(n,numObsPerChunk*(j-1) + 1); iend = min(n,numObsPerChunk*j); idx = ibegin:iend; Mdl = fit(Mdl,X(idx,:),Y(idx)); beta1(j) = Mdl.Beta(1); numtrainobs(j) = Mdl.NumTrainingObservations; priormoved(j) = Mdl.Prior(Mdl.ClassNames == true); end
Mdl is an incrementalClassificationLinear model object trained on all the data in the stream.
To see how the parameters evolve during incremental learning, plot them on separate tiles.
tiledlayout(2,2) nexttile plot(beta1) ylabel('\beta_1') xline(Mdl.EstimationPeriod/numObsPerChunk,'r-.') xlabel('Iteration') axis tight nexttile plot(numtrainobs) ylabel('Number of Training Observations') xline(Mdl.EstimationPeriod/numObsPerChunk,'r-.') xlabel('Iteration') axis tight nexttile plot(priormoved) ylabel('\pi(Subject Is Moving)') xline(Mdl.EstimationPeriod/numObsPerChunk,'r-.') xlabel('Iteration') axis tight
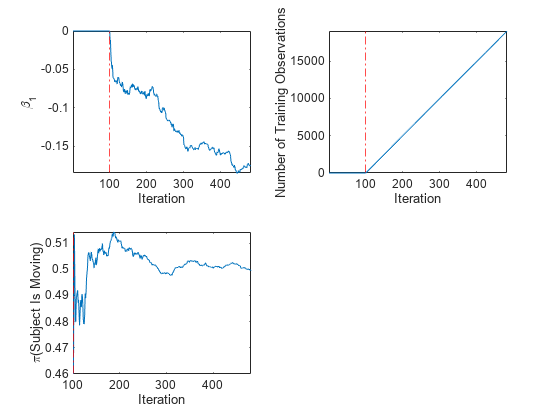
The plot suggests that fit does not fit the model to the data or update the parameters until after the estimation period.
Train a linear model for binary classification by using fitclinear, convert it to an incremental learner, track its performance, and fit it to streaming data. Orient the observations in columns, and specify observation weights.
Load and Preprocess Data
Load the human activity data set. Randomly shuffle the data. Orient the observations of the predictor data in columns.
load humanactivity rng(1); % For reproducibility n = numel(actid); idx = randsample(n,n); X = feat(idx,:)'; Y = actid(idx);
For details on the data set, enter Description at the command line.
Responses can be one of five classes: Sitting, Standing, Walking, Running, or Dancing. Dichotomize the response by identifying whether the subject is moving (actid > 2).
Y = Y > 2;
Suppose that the data collected when the subject was not moving (Y = false) has double the quality than when the subject was moving. Create a weight variable that attributes 2 to observations collected from a still subject, and 1 to a moving subject.
W = ones(n,1) + ~Y;
Train Linear Model for Binary Classification
Fit a linear model for binary classification to a random sample of half the data.
idxtt = randsample([true false],n,true); TTMdl = fitclinear(X(:,idxtt),Y(idxtt),'ObservationsIn','columns', ... 'Weights',W(idxtt))
TTMdl =
ClassificationLinear
ResponseName: 'Y'
ClassNames: [0 1]
ScoreTransform: 'none'
Beta: [60×1 double]
Bias: -0.1107
Lambda: 8.2967e-05
Learner: 'svm'
Properties, Methods
TTMdl is a ClassificationLinear model object representing a traditionally trained linear model for binary classification.
Convert Trained Model
Convert the traditionally trained classification model to a binary classification linear model for incremental learning.
IncrementalMdl = incrementalLearner(TTMdl)
IncrementalMdl =
incrementalClassificationLinear
IsWarm: 1
Metrics: [1×2 table]
ClassNames: [0 1]
ScoreTransform: 'none'
Beta: [60×1 double]
Bias: -0.1107
Learner: 'svm'
Properties, Methods
Separately Track Performance Metrics and Fit Model
Perform incremental learning on the rest of the data by using the updateMetrics and fit functions. At each iteration:
Simulate a data stream by processing 50 observations at a time.
Call
updateMetricsto update the cumulative and window classification error of the model given the incoming chunk of observations. Overwrite the previous incremental model to update the losses in theMetricsproperty. Note that the function does not fit the model to the chunk of data—the chunk is "new" data for the model. Specify that the observations are oriented in columns, and specify the observation weights.Call
fitto fit the model to the incoming chunk of observations. Overwrite the previous incremental model to update the model parameters. Specify that the observations are oriented in columns, and specify the observation weights.Store the classification error and first estimated coefficient .
% Preallocation idxil = ~idxtt; nil = sum(idxil); numObsPerChunk = 50; nchunk = floor(nil/numObsPerChunk); ce = array2table(zeros(nchunk,2),'VariableNames',["Cumulative" "Window"]); beta1 = [IncrementalMdl.Beta(1); zeros(nchunk,1)]; Xil = X(:,idxil); Yil = Y(idxil); Wil = W(idxil); % Incremental fitting for j = 1:nchunk ibegin = min(nil,numObsPerChunk*(j-1) + 1); iend = min(nil,numObsPerChunk*j); idx = ibegin:iend; IncrementalMdl = updateMetrics(IncrementalMdl,Xil(:,idx),Yil(idx), ... 'ObservationsIn','columns','Weights',Wil(idx)); ce{j,:} = IncrementalMdl.Metrics{"ClassificationError",:}; IncrementalMdl = fit(IncrementalMdl,Xil(:,idx),Yil(idx),'ObservationsIn','columns', ... 'Weights',Wil(idx)); beta1(j + 1) = IncrementalMdl.Beta(1); end
IncrementalMdl is an incrementalClassificationLinear model object trained on all the data in the stream.
Alternatively, you can use updateMetricsAndFit to update performance metrics of the model given a new chunk of data, and then fit the model to the data.
Plot a trace plot of the performance metrics and estimated coefficient .
t = tiledlayout(2,1); nexttile h = plot(ce.Variables); xlim([0 nchunk]) ylabel('Classification Error') legend(h,ce.Properties.VariableNames) nexttile plot(beta1) ylabel('\beta_1') xlim([0 nchunk]) xlabel(t,'Iteration')

The cumulative loss is stable and gradually decreases, whereas the window loss jumps.
changes gradually, then levels off, as fit processes more chunks.
Incrementally train a linear regression model only when its performance degrades.
Load and shuffle the 2015 NYC housing data set. For more details on the data, see NYC Open Data.
load NYCHousing2015 rng(1) % For reproducibility n = size(NYCHousing2015,1); shuffidx = randsample(n,n); NYCHousing2015 = NYCHousing2015(shuffidx,:);
Extract the response variable SALEPRICE from the table. For numerical stability, scale SALEPRICE by 1e6.
Y = NYCHousing2015.SALEPRICE/1e6; NYCHousing2015.SALEPRICE = [];
Create dummy variable matrices from the categorical predictors.
catvars = ["BOROUGH" "BUILDINGCLASSCATEGORY" "NEIGHBORHOOD"]; dumvarstbl = varfun(@(x)dummyvar(categorical(x)),NYCHousing2015, ... 'InputVariables',catvars); dumvarmat = table2array(dumvarstbl); NYCHousing2015(:,catvars) = [];
Treat all other numeric variables in the table as linear predictors of sales price. Concatenate the matrix of dummy variables to the rest of the predictor data.
idxnum = varfun(@isnumeric,NYCHousing2015,'OutputFormat','uniform'); X = [dumvarmat NYCHousing2015{:,idxnum}];
Configure a linear regression model for incremental learning so that it does not have an estimation or metrics warm-up period. Specify a metrics window size of 1000. Fit the configured model to the first 100 observations.
Mdl = incrementalRegressionLinear('EstimationPeriod',0, ... 'MetricsWarmupPeriod',0,'MetricsWindowSize',1000); numObsPerChunk = 100; Mdl = fit(Mdl,X(1:numObsPerChunk,:),Y(1:numObsPerChunk));
Mdl is an incrementalRegressionLinear model object.
Perform incremental learning, with conditional fitting, by following this procedure for each iteration:
Simulate a data stream by processing a chunk of 100 observations at a time.
Update the model performance by computing the epsilon insensitive loss, within a 200 observation window.
Fit the model to the chunk of data only when the loss more than doubles from the minimum loss experienced.
When tracking performance and fitting, overwrite the previous incremental model.
Store the epsilon insensitive loss and to see how the loss and coefficient evolve during training.
Track when
fittrains the model.
% Preallocation n = numel(Y) - numObsPerChunk; nchunk = floor(n/numObsPerChunk); beta313 = zeros(nchunk,1); ei = array2table(nan(nchunk,2),'VariableNames',["Cumulative" "Window"]); trained = false(nchunk,1); % Incremental fitting for j = 2:nchunk ibegin = min(n,numObsPerChunk*(j-1) + 1); iend = min(n,numObsPerChunk*j); idx = ibegin:iend; Mdl = updateMetrics(Mdl,X(idx,:),Y(idx)); ei{j,:} = Mdl.Metrics{"EpsilonInsensitiveLoss",:}; minei = min(ei{:,2}); pdiffloss = (ei{j,2} - minei)/minei*100; if pdiffloss > 100 Mdl = fit(Mdl,X(idx,:),Y(idx)); trained(j) = true; end beta313(j) = Mdl.Beta(end); end
Mdl is an incrementalRegressionLinear model object trained on all the data in the stream.
To see how the model performance and evolve during training, plot them on separate tiles.
t = tiledlayout(2,1); nexttile plot(beta313) hold on plot(find(trained),beta313(trained),'r.') xlim([0 nchunk]) ylabel('\beta_{313}') xline(Mdl.EstimationPeriod/numObsPerChunk,'r-.') legend('\beta_{313}','Training occurs','Location','southeast') hold off nexttile plot(ei.Variables) xlim([0 nchunk]) ylabel('Epsilon Insensitive Loss') xline(Mdl.EstimationPeriod/numObsPerChunk,'r-.') legend(ei.Properties.VariableNames) xlabel(t,'Iteration')

The trace plot of shows periods of constant values, during which the loss did not double from the minimum experienced.
Input Arguments
Incremental learning model to fit to streaming data, specified as an incrementalClassificationLinear or incrementalRegressionLinear model object. You can create
Mdl directly or by converting a supported, traditionally trained
machine learning model using the incrementalLearner function. For
more details, see the corresponding reference page.
Chunk of predictor data to which the model is fit, specified as a floating-point
matrix of n observations and Mdl.NumPredictors
predictor variables. The value of the ObservationsIn name-value
argument determines the orientation of the variables and observations. The default
ObservationsIn value is "rows", which indicates that
observations in the predictor data are oriented along the rows of
X.
The length of the observation labels Y and the number of
observations in X must be equal;
Y( is the label of observation
j (row or column) in j)X.
Note
If
Mdl.NumPredictors= 0,fitinfers the number of predictors fromX, and sets the corresponding property of the output model. Otherwise, if the number of predictor variables in the streaming data changes fromMdl.NumPredictors,fitissues an error.fitsupports only floating-point input predictor data. If your input data includes categorical data, you must prepare an encoded version of the categorical data. Usedummyvarto convert each categorical variable to a numeric matrix of dummy variables. Then, concatenate all dummy variable matrices and any other numeric predictors. For more details, see Dummy Variables.
Data Types: single | double
Chunk of responses (labels) to which the model is fit, specified as a categorical, character, or string array, logical or floating-point vector, or cell array of character vectors for classification problems; or a floating-point vector for regression problems.
The length of the observation labels Y and the number of observations in X must be equal; Y( is the label of observation j (row or column) in j)X.
For classification problems:
fitsupports binary classification only.When the
ClassNamesproperty of the input modelMdlis nonempty, the following conditions apply:If
Ycontains a label that is not a member ofMdl.ClassNames,fitissues an error.The data type of
YandMdl.ClassNamesmust be the same.
Data Types: char | string | cell | categorical | logical | single | double
Note
If an observation (predictor or label) or weight contains at least one missing (
NaN) value,fitignores the observation. Consequently,fituses fewer than n observations to create an updated model, where n is the number of observations inX.The chunk size n and the stochastic gradient descent (SGD) hyperparameter mini-batch size (
Mdl.BatchSize) can be different values, and n does not have to be an exact multiple of the mini-batch size. If n <Mdl.BatchSize,fituses the n available observations when it applies SGD. If n >Mdl.BatchSize, the function updates the model with a mini-batch of the specified size multiple times, and then uses the rest of the observations for the last mini-batch. The number of observations for the last mini-batch can be smaller thanMdl.BatchSize.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: 'ObservationsIn','columns','Weights',W specifies that the
columns of the predictor matrix correspond to observations, and the vector
W contains observation weights to apply during incremental
learning.
Predictor data observation dimension, specified as the comma-separated pair consisting of 'ObservationsIn' and 'columns' or 'rows'.
Data Types: char | string
Chunk of observation weights, specified as the comma-separated pair consisting of 'Weights' and a floating-point vector of positive values. fit weighs the observations in X with the corresponding values in Weights. The size of Weights must equal n, which is the number of observations in X.
By default, Weights is ones(.n,1)
For more details, including normalization schemes, see Observation Weights.
Data Types: double | single
Output Arguments
Updated incremental learning model, returned as an incremental learning model object of the same data type as the input model Mdl, either incrementalClassificationLinear or incrementalRegressionLinear.
If Mdl.EstimationPeriod > 0, the incremental fitting functions
updateMetricsAndFit and fit estimate
hyperparameters using the first Mdl.EstimationPeriod observations
passed to either function; they do not train the input model to that data. However, if
an incoming chunk of n observations is greater than or equal to the
number of observations remaining in the estimation period m,
fit estimates hyperparameters using the first
n – m observations, and fits the input model to
the remaining m observations. Consequently, the software updates the
Beta and Bias properties, hyperparameter
properties, and recordkeeping properties such as
NumTrainingObservations.
For classification problems, if the ClassNames property of the input model Mdl is an empty array, fit sets the ClassNames property of the output model Mdl to unique(Y).
Tips
Unlike traditional training, incremental learning might not have a separate test (holdout) set. Therefore, to treat each incoming chunk of data as a test set, pass the incremental model and each incoming chunk to
updateMetricsbefore training the model on the same data.
Algorithms
For classification problems, if the prior class probability distribution is known (in other words, the prior distribution is not empirical), fit normalizes observation weights to sum to the prior class probabilities in the respective classes. This action implies that observation weights are the respective prior class probabilities by default.
For regression problems or if the prior class probability distribution is empirical, the software normalizes the specified observation weights to sum to 1 each time you call fit.
Extended Capabilities
Usage notes and limitations:
Use
saveLearnerForCoder,loadLearnerForCoder, andcodegen(MATLAB Coder) to generate code for thefitfunction. Save a trained model by usingsaveLearnerForCoder. Define an entry-point function that loads the saved model by usingloadLearnerForCoderand calls thefitfunction. Then usecodegento generate code for the entry-point function.To generate single-precision C/C++ code for
fit, specifyDataType="single"when you call theloadLearnerForCoderfunction.This table contains notes about the arguments of
fit. Arguments not included in this table are fully supported.Argument Notes and Limitations MdlFor usage notes and limitations of the model object, see
incrementalClassificationLinearorincrementalRegressionLinear.XBatch-to-batch, the number of observations can be a variable size, but must equal the number of observations in
Y.The number of predictor variables must equal
Mdl.NumPredictors.Xmust besingleordouble.
YBatch-to-batch, the number of observations can be a variable size, but must equal the number of observations in
X.For classification problems, all labels in
Ymust be represented inMdl.ClassNames.YandMdl.ClassNamesmust have the same data type.
If you configure
Mdlto shuffle data (Mdl.Shuffleistrue, orMdl.Solveris"sgd"or"asgd"), thefitfunction randomly shuffles each incoming batch of observations before it fits the model to the batch. The order of the shuffled observations might not match the order generated by MATLAB®. Therefore, the fitted coefficients computed in MATLAB might not be equal to those computed by the generated code.Use a homogeneous data type (specifically,
singleordouble) for all floating-point input arguments and object properties.
For more information, see Introduction to Code Generation for Statistics and Machine Learning Functions.
Version History
Introduced in R2020b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleziona un sito web
Seleziona un sito web per visualizzare contenuto tradotto dove disponibile e vedere eventi e offerte locali. In base alla tua area geografica, ti consigliamo di selezionare: .
Puoi anche selezionare un sito web dal seguente elenco:
Come ottenere le migliori prestazioni del sito
Per ottenere le migliori prestazioni del sito, seleziona il sito cinese (in cinese o in inglese). I siti MathWorks per gli altri paesi non sono ottimizzati per essere visitati dalla tua area geografica.
Americhe
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)