cvloss
Description
L = cvloss(CVMdl)CVMdl at each step of the horizon (CVMdl.Horizon).
For each partition window in CVMdl.Partition and each horizon step, the
function computes the loss for the test observations using a model trained on the training
observations. CVMdl.X and CVMdl.Y contain all
observations.
L = cvloss(CVMdl,Name=Value)
Examples
Create a cross-validated direct forecasting model using expanding window cross-validation. To evaluate the performance of the model:
Compute the mean squared error (MSE) on each test set using the
cvlossobject function.For each test set, compare the true response values to the predicted response values using the
cvpredictobject function.
Load the sample file TemperatureData.csv, which contains average daily temperature from January 2015 through July 2016. Read the file into a table. Observe the first eight observations in the table.
Tbl = readtable("TemperatureData.csv");
head(Tbl) Year Month Day TemperatureF
____ ___________ ___ ____________
2015 {'January'} 1 23
2015 {'January'} 2 31
2015 {'January'} 3 25
2015 {'January'} 4 39
2015 {'January'} 5 29
2015 {'January'} 6 12
2015 {'January'} 7 10
2015 {'January'} 8 4
Create a datetime variable t that contains the year, month, and day information for each observation in Tbl.
numericMonth = month(datetime(Tbl.Month, ... InputFormat="MMMM",Locale="en_US")); t = datetime(Tbl.Year,numericMonth,Tbl.Day);
Plot the temperature values in Tbl over time.
plot(t,Tbl.TemperatureF) xlabel("Date") ylabel("Temperature in Fahrenheit")
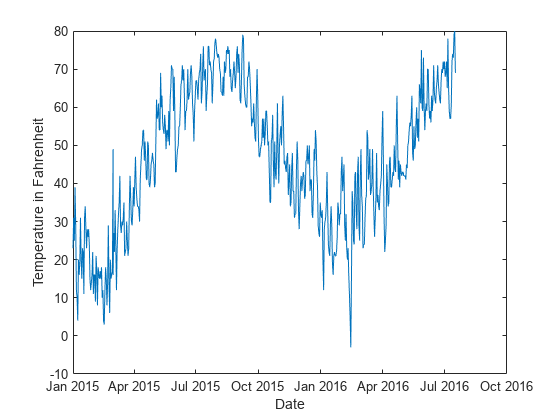
Create a direct forecasting model by using the data in Tbl. Train the model using a bagged ensemble of trees. All three of the predictors (Year, Month, and Day) are leading predictors because their future values are known. To create new predictors by shifting the leading predictor and response variables backward in time, specify the leading predictor lags and the response variable lags.
Mdl = directforecaster(Tbl,"TemperatureF", ... Learner="bag", ... LeadingPredictors="all",LeadingPredictorLags={0:1,0:1,0:7}, ... ResponseLags=1:7)
Mdl =
DirectForecaster
Horizon: 1
ResponseLags: [1 2 3 4 5 6 7]
LeadingPredictors: [1 2 3]
LeadingPredictorLags: {[0 1] [0 1] [0 1 2 3 4 5 6 7]}
ResponseName: 'TemperatureF'
PredictorNames: {'Year' 'Month' 'Day'}
CategoricalPredictors: 2
Learners: {[1×1 classreg.learning.regr.CompactRegressionEnsemble]}
MaxLag: 7
NumObservations: 565
Properties, Methods
Mdl is a DirectForecaster model object. By default, the horizon is one step ahead. That is, Mdl predicts a value that is one step into the future.
Partition the time series data in Tbl using an expanding window cross-validation scheme. Create three training sets and three test sets, where each test set has 100 observations. Note that each observation in Tbl is in at most one test set.
CVPartition = tspartition(size(Mdl.X,1),"ExpandingWindow",3, ... TestSize=100)
CVPartition =
tspartition
Type: 'expanding-window'
NumObservations: 565
NumTestSets: 3
TrainSize: [265 365 465]
TestSize: [100 100 100]
StepSize: 100
Properties, Methods
The training sets increase in size from 265 observations in the first window to 465 observations in the third window.
Create a cross-validated direct forecasting model using the partition specified in CVPartition. Inspect the Learners property of the resulting CVMdl object.
CVMdl = crossval(Mdl,CVPartition)
CVMdl =
PartitionedDirectForecaster
Partition: [1×1 tspartition]
Horizon: 1
ResponseLags: [1 2 3 4 5 6 7]
LeadingPredictors: [1 2 3]
LeadingPredictorLags: {[0 1] [0 1] [0 1 2 3 4 5 6 7]}
ResponseName: 'TemperatureF'
PredictorNames: {'Year' 'Month' 'Day'}
CategoricalPredictors: 2
Learners: {3×1 cell}
MaxLag: 7
NumObservations: 565
Properties, Methods
CVMdl.Learners
ans=3×1 cell array
{1×1 timeseries.forecaster.CompactDirectForecaster}
{1×1 timeseries.forecaster.CompactDirectForecaster}
{1×1 timeseries.forecaster.CompactDirectForecaster}
CVMdl is a PartitionedDirectForecaster model object. The crossval function trains CVMdl.Learners{1} using the observations in the first training set, CVMdl.Learner{2} using the observations in the second training set, and CVMdl.Learner{3} using the observations in the third training set.
Compute the average test set MSE.
averageMSE = cvloss(CVMdl)
averageMSE = 53.3480
To obtain more information, compute the MSE for each test set.
individualMSE = cvloss(CVMdl,Mode="individual")individualMSE = 3×1
44.1352
84.0695
31.8393
The models trained on the first and third training sets seem to perform better than the model trained on the second training set.
For each test set observation, predict the temperature value using the corresponding model in CVMdl.Learners.
predictedY = cvpredict(CVMdl); predictedY(260:end,:)
ans=306×1 table
TemperatureF_Step1
__________________
NaN
NaN
NaN
NaN
NaN
NaN
50.963
57.363
57.04
60.705
59.606
58.302
58.023
61.39
67.229
61.083
⋮
Only the last 300 observations appear in any test set. For observations that do not appear in a test set, the predicted response value is NaN.
For each test set, plot the true response values and the predicted response values.
tiledlayout(3,1) nexttile idx1 = test(CVPartition,1); plot(t(idx1),Tbl.TemperatureF(idx1)) hold on plot(t(idx1),predictedY.TemperatureF_Step1(idx1)) legend("True Response","Predicted Response", ... Location="eastoutside") xlabel("Date") ylabel("Temperature") title("Test Set 1") hold off nexttile idx2 = test(CVPartition,2); plot(t(idx2),Tbl.TemperatureF(idx2)) hold on plot(t(idx2),predictedY.TemperatureF_Step1(idx2)) legend("True Response","Predicted Response", ... Location="eastoutside") xlabel("Date") ylabel("Temperature") title("Test Set 2") hold off nexttile idx3 = test(CVPartition,3); plot(t(idx3),Tbl.TemperatureF(idx3)) hold on plot(t(idx3),predictedY.TemperatureF_Step1(idx3)) legend("True Response","Predicted Response", ... Location="eastoutside") xlabel("Date") ylabel("Temperature") title("Test Set 3") hold off
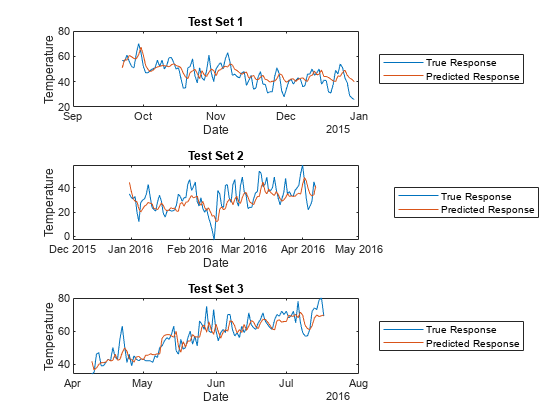
Overall, the cross-validated direct forecasting model is able to predict the trend in temperatures. If you are satisfied with the performance of the cross-validated model, you can use the full DirectForecaster model Mdl for forecasting at time steps beyond the available data.
Create a partitioned direct forecasting model using holdout validation. To evaluate the performance of the model:
At each horizon step, compute the root relative squared error (RRSE) on the test set using the
cvlossobject function.At each horizon step, compare the true response values to the predicted response values using the
cvpredictobject function.
Load the sample file TemperatureData.csv, which contains average daily temperature from January 2015 through July 2016. Read the file into a table. Observe the first eight observations in the table.
Tbl = readtable("TemperatureData.csv");
head(Tbl) Year Month Day TemperatureF
____ ___________ ___ ____________
2015 {'January'} 1 23
2015 {'January'} 2 31
2015 {'January'} 3 25
2015 {'January'} 4 39
2015 {'January'} 5 29
2015 {'January'} 6 12
2015 {'January'} 7 10
2015 {'January'} 8 4
Create a datetime variable t that contains the year, month, and day information for each observation in Tbl.
numericMonth = month(datetime(Tbl.Month, ... InputFormat="MMMM",Locale="en_US")); t = datetime(Tbl.Year,numericMonth,Tbl.Day);
Plot the temperature values in Tbl over time.
plot(t,Tbl.TemperatureF) xlabel("Date") ylabel("Temperature in Fahrenheit")
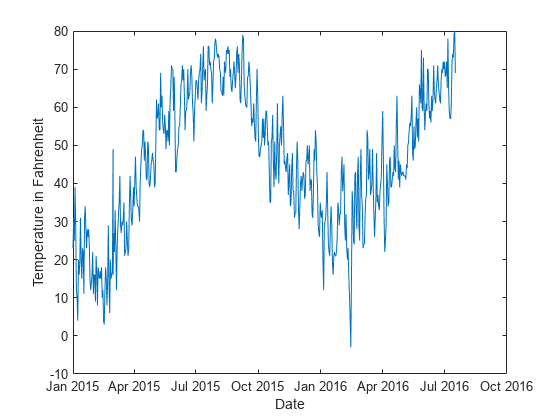
Create a direct forecasting model by using the data in Tbl. Specify the horizon steps as one, two, and three steps ahead. Train a model at each horizon using a bagged ensemble of trees. All three of the predictors (Year, Month, and Day) are leading predictors because their future values are known. To create new predictors by shifting the leading predictor and response variables backward in time, specify the leading predictor lags and the response variable lags.
rng("default") Mdl = directforecaster(Tbl,"TemperatureF", ... Horizon=1:3,Learner="bag", ... LeadingPredictors="all",LeadingPredictorLags={0:1,0:1,0:7}, ... ResponseLags=1:7)
Mdl =
DirectForecaster
Horizon: [1 2 3]
ResponseLags: [1 2 3 4 5 6 7]
LeadingPredictors: [1 2 3]
LeadingPredictorLags: {[0 1] [0 1] [0 1 2 3 4 5 6 7]}
ResponseName: 'TemperatureF'
PredictorNames: {'Year' 'Month' 'Day'}
CategoricalPredictors: 2
Learners: {3×1 cell}
MaxLag: 7
NumObservations: 565
Properties, Methods
Mdl is a DirectForecaster model object. Mdl consists of three regression models: Mdl.Learners{1}, which predicts one step ahead; Mdl.Learners{2}, which predicts two steps ahead; and Mdl.Learners{3}, which predicts three steps ahead.
Partition the time series data in Tbl using a holdout validation scheme. Reserve 20% of the observations for testing.
holdoutPartition = tspartition(size(Mdl.X,1),"Holdout",0.20)holdoutPartition =
tspartition
Type: 'holdout'
NumObservations: 565
NumTestSets: 1
TrainSize: 452
TestSize: 113
Properties, Methods
The test set consists of the latest 113 observations.
Create a partitioned direct forecasting model using the partition specified in holdoutPartition.
holdoutMdl = crossval(Mdl,holdoutPartition)
holdoutMdl =
PartitionedDirectForecaster
Partition: [1×1 tspartition]
Horizon: [1 2 3]
ResponseLags: [1 2 3 4 5 6 7]
LeadingPredictors: [1 2 3]
LeadingPredictorLags: {[0 1] [0 1] [0 1 2 3 4 5 6 7]}
ResponseName: 'TemperatureF'
PredictorNames: {'Year' 'Month' 'Day'}
CategoricalPredictors: 2
Learners: {[1×1 timeseries.forecaster.CompactDirectForecaster]}
MaxLag: 7
NumObservations: 565
Properties, Methods
holdoutMdl is a PartitionedDirectForecaster model object. Because holdoutMdl uses holdout validation rather than a cross-validation scheme, the Learners property of the object contains one CompactDirectForecaster model only.
Like Mdl, holdoutMdl contains three regression models. The crossval function trains holdoutMdl.Learners{1}.Learners{1}, holdoutMdl.Learners{1}.Learners{2}, and holdoutMdl.Learners{1}.Learners{3} using the same training data. However, the three models use different response variables because each model predicts values for a different horizon step.
holdoutMdl.Learners{1}.Learners{1}.ResponseNameans = 'TemperatureF_Step1'
holdoutMdl.Learners{1}.Learners{2}.ResponseNameans = 'TemperatureF_Step2'
holdoutMdl.Learners{1}.Learners{3}.ResponseNameans = 'TemperatureF_Step3'
Compute the root relative squared error (RRSE) on the test data at each horizon step. Use the helper function computeRRSE (shown at the end of this example). The RRSE indicates how well a model performs relative to the simple model, which always predicts the average of the true values. In particular, when the RRSE is less than 1, the model performs better than the simple model.
holdoutRRSE = cvloss(holdoutMdl,LossFun=@computeRRSE)
holdoutRRSE = 1×3
0.4797 0.5889 0.6103
At each horizon, the direct forecasting model seems to perform better than the simple model.
For each test set observation, predict the temperature value using the corresponding model in holdoutMdl.Learners.
predictedY = cvpredict(holdoutMdl); predictedY(450:end,:)
ans=116×3 table
TemperatureF_Step1 TemperatureF_Step2 TemperatureF_Step3
__________________ __________________ __________________
NaN NaN NaN
NaN NaN NaN
NaN NaN NaN
41.063 39.758 41.234
33.721 36.507 37.719
36.987 35.133 37.719
38.644 34.598 36.444
38.917 34.576 36.275
45.888 37.005 38.34
48.516 42.762 41.05
44.882 46.816 43.881
35.057 45.301 47.048
31.1 41.473 42.948
31.817 37.314 42.946
33.166 38.419 41.3
40.279 38.432 40.533
⋮
Recall that only the latest 113 observations appear in the test set. For observations that do not appear in the test set, the predicted response value is NaN.
For each test set, plot the true response values and the predicted response values.
tiledlayout(3,1) idx = test(holdoutPartition); nexttile plot(t(idx),Tbl.TemperatureF(idx)) hold on plot(t(idx),predictedY.TemperatureF_Step1(idx)) legend("True Response","Predicted Response", ... Location="eastoutside") xlabel("Date") ylabel("Temperature") title("Horizon 1") hold off nexttile plot(t(idx),Tbl.TemperatureF(idx)) hold on plot(t(idx),predictedY.TemperatureF_Step2(idx)) legend("True Response","Predicted Response", ... Location="eastoutside") xlabel("Date") ylabel("Temperature") title("Horizon 2") hold off nexttile plot(t(idx),Tbl.TemperatureF(idx)) hold on plot(t(idx),predictedY.TemperatureF_Step3(idx)) legend("True Response","Predicted Response", ... Location="eastoutside") xlabel("Date") ylabel("Temperature") title("Horizon 3") hold off

Overall, holdoutMdl is able to predict the trend in temperatures, although it seems to perform best when forecasting one step ahead. If you are satisfied with the performance of the partitioned model, you can use the full DirectForecaster model Mdl for forecasting at time steps beyond the available data.
Helper Function
The helper function computeRRSE computes the RRSE given the true response variable trueY and the predicted values predY. This code creates the computeRRSE helper function.
function rrse = computeRRSE(trueY,predY) error = trueY(:) - predY(:); meanY = mean(trueY(:),"omitnan"); rrse = sqrt(sum(error.^2,"omitnan")/sum((trueY(:) - meanY).^2,"omitnan")); end
Input Arguments
Cross-validated direct forecasting model, specified as a PartitionedDirectForecaster model object.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: cvloss(CVMdl,Mode="individual") specifies to return the loss
for each window and horizon step combination.
Loss function, specified as "mse" or a function handle.
If you specify the built-in function
"mse", then the loss function is the mean squared error.If you specify your own function using function handle notation, then the function must have the signature
lossvalue =, where:lossfun(Y,predictedY)The output argument
lossvalueis a scalar.You specify the function name (
lossfun).Yis an n-by-1 vector of observed numeric responses at a specific horizon step, where n is the number of observations (CVMdl.NumObservations).predictedYis an n-by-1 vector of predicted numeric responses at a specific horizon step.
Specify your function using
LossFun=@.lossfun
Data Types: single | double | function_handle
Loss aggregation level, specified as "average" or
"individual".
| Value | Description |
|---|---|
"average" |
At each horizon step, if an observation is in more than one test set, the function averages the predictions for the observation over all test sets before computing the loss. |
"individual" |
Before computing loss values at each horizon step, the function does not average the predictions for observations that are in more than one test set. |
Example: Model="individual"
Data Types: char | string
Output Arguments
Losses, returned as a numeric vector or numeric matrix.
If
Modeis"average", thenLis a vector of loss values containing the loss at each horizon step, averaged over all partition windows. At each horizon step, if an observation is in more than one test set, the function averages the predictions for the observation over all test sets before computing the loss.If
Modeis"individual", thenLis a w-by-h matrix of loss values, where w is the number of partition windows and h is the number of horizon steps (that is, the number of elements inCVMdl.Horizon). Before computing the loss values at each horizon step, the function does not average the predictions for observations that are in more than one test set.
Version History
Introduced in R2023b
See Also
PartitionedDirectForecaster | cvpredict | DirectForecaster | tspartition
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleziona un sito web
Seleziona un sito web per visualizzare contenuto tradotto dove disponibile e vedere eventi e offerte locali. In base alla tua area geografica, ti consigliamo di selezionare: .
Puoi anche selezionare un sito web dal seguente elenco:
Come ottenere le migliori prestazioni del sito
Per ottenere le migliori prestazioni del sito, seleziona il sito cinese (in cinese o in inglese). I siti MathWorks per gli altri paesi non sono ottimizzati per essere visitati dalla tua area geografica.
Americhe
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)