plot
Visualize prior and posterior densities of Bayesian linear regression model parameters
Syntax
Description
plot( or
PosteriorMdl)plot( plots the posterior or
prior distributions of the parameters in the Bayesian linear regression
model
PriorMdl)PosteriorMdl or PriorMdl, respectively.
plot adds subplots for each parameter to one figure and
overwrites the same figure when you call plot multiple
times.
plot(
plots the posterior and prior distributions in the same subplot.
PosteriorMdl,PriorMdl)plot uses solid blue lines for posterior
densities and dashed red lines for prior densities.
plot(___,
uses any of the input argument combinations in the previous syntaxes and
additional options specified by one or more name-value pair arguments. For
example, you can evaluate the posterior or prior density by supplying values of
β and σ2,
or choose which parameter distributions to include in the figure.Name,Value)
pointsUsed = plot(___)plot
uses to evaluate the densities in the subplots.
[
also returns the values of the evaluated densities.pointsUsed,posteriorDensity,priorDensity]
= plot(___)
If you specify one model, then plot returns the
density values in PosteriorDensity. Otherwise,
plot returns the posterior density values in
PosteriorDensity and the prior density values in
PriorDensity.
[
returns the figure handle of the figure containing the distributions.pointsUsed,posteriorDensity,priorDensity,FigureHandle]
= plot(___)
Examples
Consider the multiple linear regression model that predicts the US real gross national product (GNPR) using a linear combination of industrial production index (IPI), total employment (E), and real wages (WR).
For all , is a series of independent Gaussian disturbances with a mean of 0 and variance .
Assume these prior distributions:
. is a 4-by-1 vector of means, and is a scaled 4-by-4 positive definite covariance matrix.
. and are the shape and scale, respectively, of an inverse gamma distribution.
These assumptions and the data likelihood imply a normal-inverse-gamma conjugate model.
Create a normal-inverse-gamma conjugate prior model for the linear regression parameters. Specify the number of predictors p and the variable names.
p = 3; VarNames = ["IPI" "E" "WR"]; PriorMdl = bayeslm(p,'ModelType','conjugate','VarNames',VarNames);
PriorMdl is a conjugateblm Bayesian linear regression model object representing the prior distribution of the regression coefficients and disturbance variance.
Plot the prior distributions.
plot(PriorMdl);

plot plots the marginal prior distributions of the intercept, regression coefficients, and disturbance variance.
Suppose that the mean of the regression coefficients is and their scaled covariance matrix is
Also, the prior scale of the disturbance variance is 0.01. Specify the prior information using dot notation.
PriorMdl.Mu = [-20; 4; 0.001; 2]; PriorMdl.V = diag([1 0.001 1e-8 0.01]); PriorMdl.B = 0.01;
Request a new figure and plot the prior distribution.
plot(PriorMdl);
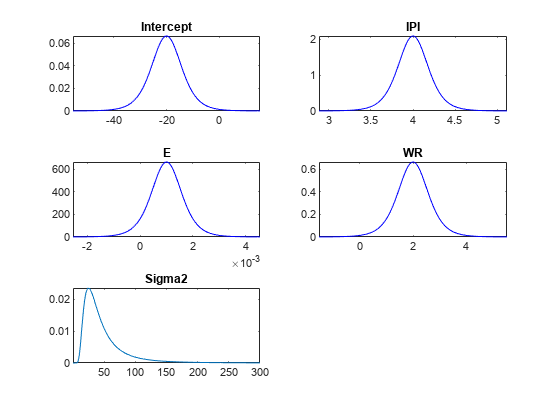
plot replaces the current distribution figure with a plot of the prior distribution of the disturbance variance.
Load the Nelson-Plosser data set, and create variables for the predictor and response data.
load Data_NelsonPlosser
X = DataTable{:,PriorMdl.VarNames(2:end)};
y = DataTable.GNPR;Estimate the posterior distributions.
PosteriorMdl = estimate(PriorMdl,X,y,'Display',false);PosteriorMdl is a conjugateblm model object that contains the posterior distributions of and .
Plot the posterior distributions.
plot(PosteriorMdl);

Plot the prior and posterior distributions of the parameters on the same subplots.
plot(PosteriorMdl,PriorMdl);
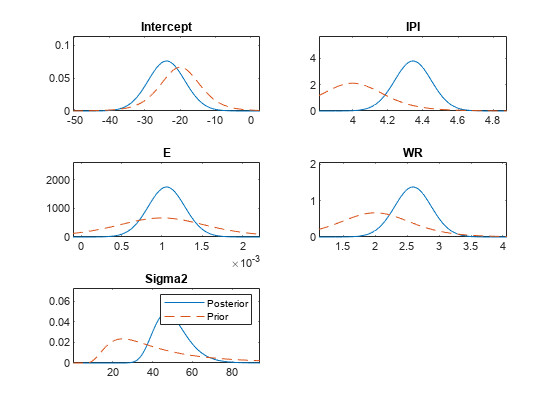
Consider the regression model in Plot Prior and Posterior Distributions.
Load the Nelson-Plosser data set, create a default conjugate prior model, and then estimate the posterior using the first 75% of the data. Turn off the estimation display.
p = 3; VarNames = ["IPI" "E" "WR"]; PriorMdl = bayeslm(p,'ModelType','conjugate','VarNames',VarNames); load Data_NelsonPlosser X = DataTable{:,PriorMdl.VarNames(2:end)}; y = DataTable.GNPR; d = 0.75; PosteriorMdlFirst = estimate(PriorMdl,X(1:floor(d*end),:),y(1:floor(d*end)),... 'Display',false);
Plot the prior distribution and the posterior distribution of the disturbance variance. Return the figure handle.
[~,~,~,h] = plot(PosteriorMdlFirst,PriorMdl,'VarNames','Sigma2');

h is the figure handle for the distribution plot. If you change the tag name of the figure by changing the Tag property, then the next plot call places all new distribution plots on a different figure.
Change the name of the figure handle to FirstHalfData using dot notation.
h.Tag = 'FirstHalfData';Estimate the posterior distribution using the rest of the data. Specify the posterior distribution based on the final 25% of the data as the prior distribution.
PosteriorMdl = estimate(PosteriorMdlFirst,X(ceil(d*end):end,:),... y(ceil(d*end):end),'Display',false);
Plot the posterior of the disturbance variance based on half of the data and all the data to a new figure.
plot(PosteriorMdl,PosteriorMdlFirst,'VarNames','Sigma2');

This type of plot shows the evolution of the posterior distribution when you incorporate new data.
Consider the regression model in Plot Prior and Posterior Distributions.
Load the Nelson-Plosser data set and create a default conjugate prior model.
p = 3; VarNames = ["IPI" "E" "WR"]; PriorMdl = bayeslm(p,'ModelType','conjugate','VarNames',VarNames); load Data_NelsonPlosser X = DataTable{:,PriorMdl.VarNames(2:end)}; y = DataTable.GNPR;
Plot the prior distributions. Request the values of the parameters used to create the plots and their respective densities.
[pointsUsedPrior,priorDensities1] = plot(PriorMdl);

pointsUsedPrior is a 5-by-1 cell array of 1-by-1000 numeric vectors representing the values of the parameters that plot uses to plot the corresponding densities. The first element corresponds to the intercept, the next three elements correspond to the regression coefficients, and the last element corresponds to the disturbance variance. priorDensities1 has the same dimensions as pointsUsed and contains the corresponding density values.
Estimate the posterior distribution. Turn off the estimation display.
PosteriorMdl = estimate(PriorMdl,X,y,'Display',false);Plot the posterior distributions. Request the values of the parameters used to create the plots and their respective densities.
[pointsUsedPost,posteriorDensities1] = plot(PosteriorMdl);
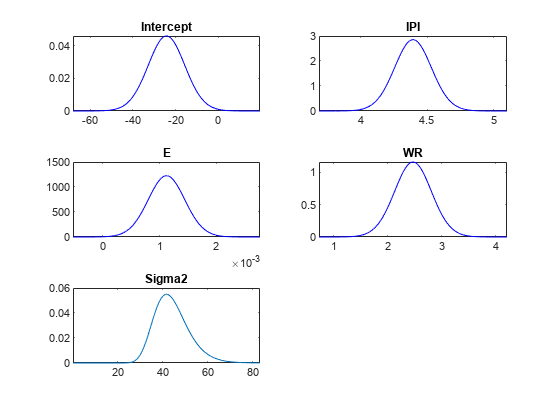
pointsUsedPost and posteriorDensities1 have the same dimensions as pointsUsedPrior. The pointsUsedPost output can be different from pointsUsedPrior. posteriorDensities1 contains the posterior density values.
Plot the prior and posterior distributions. Request the values of the parameters used to create the plots and their respective densities.
[pointsUsedPP,posteriorDensities2,priorDensities2] = plot(PosteriorMdl,PriorMdl);
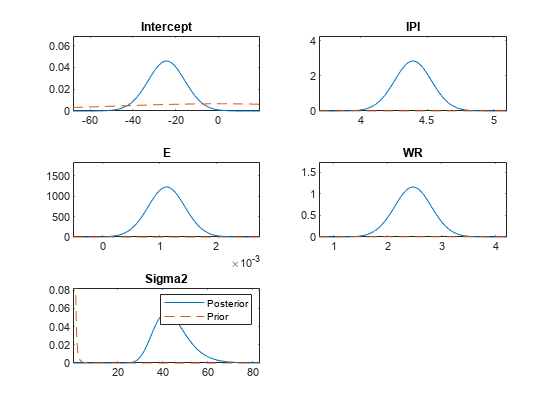
All output values have the same dimensions as pointsUsedPrior. The posteriorDensities2 output contains the posterior density values. The priorDensities2 output contains the prior density values.
Confirm that pointsUsedPP is equal to pointsUsedPost.
compare = @(a,b)sum(a == b) == numel(a); cellfun(compare,pointsUsedPost,pointsUsedPP)
ans = 5×1 logical array
1
1
1
1
1
The points used are equivalent.
Confirm that the posterior densities are the same, but that the prior densities are not.
cellfun(compare,posteriorDensities1,posteriorDensities2)
ans = 5×1 logical array
1
1
1
1
1
cellfun(compare,priorDensities1,priorDensities2)
ans = 5×1 logical array
0
0
0
0
0
When plotting only the prior distribution, plot evaluates the prior densities at points that produce a clear plot of the prior distribution. When plotting both a prior and posterior distribution, plot prefers to plot the posterior clearly. Therefore, plot can determine a different set of points to use.
Consider the regression model in Plot Prior and Posterior Distributions.
Load the Nelson-Plosser data set and create a default conjugate prior model for the regression coefficients and disturbance variance. Then, estimate the posterior distribution and obtain the estimation summary table from summarize.
p = 3; VarNames = ["IPI" "E" "WR"]; PriorMdl = bayeslm(p,'ModelType','conjugate','VarNames',VarNames); load Data_NelsonPlosser X = DataTable{:,PriorMdl.VarNames(2:end)}; y = DataTable.GNPR; PosteriorMdl = estimate(PriorMdl,X,y);
Method: Analytic posterior distributions
Number of observations: 62
Number of predictors: 4
Log marginal likelihood: -259.348
| Mean Std CI95 Positive Distribution
-----------------------------------------------------------------------------------
Intercept | -24.2494 8.7821 [-41.514, -6.985] 0.003 t (-24.25, 8.65^2, 68)
IPI | 4.3913 0.1414 [ 4.113, 4.669] 1.000 t (4.39, 0.14^2, 68)
E | 0.0011 0.0003 [ 0.000, 0.002] 1.000 t (0.00, 0.00^2, 68)
WR | 2.4683 0.3490 [ 1.782, 3.154] 1.000 t (2.47, 0.34^2, 68)
Sigma2 | 44.1347 7.8020 [31.427, 61.855] 1.000 IG(34.00, 0.00069)
summaryTbl = summarize(PosteriorMdl); summaryTbl = summaryTbl.MarginalDistributions;
summaryTbl is a table containing the statistics that estimate displays at the command line.
For each parameter, determine a set of 50 evenly spaced values within three standard deviations of the mean. Put the values into the cells of a 5-by-1 cell vector following the order of the parameters that comprise the rows of the estimation summary table.
Points = cell(numel(summaryTbl.Mean),1); % Preallocation for j = 1:numel(summaryTbl.Mean) Points{j} = linspace(summaryTbl.Mean(j) - 3*summaryTbl.Std(j),... summaryTbl.Mean(j) + 2*summaryTbl.Std(j),50); end
Plot the posterior distributions within their respective intervals.
plot(PosteriorMdl,'Points',Points)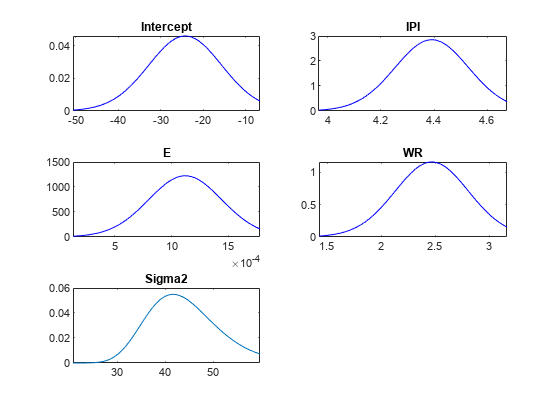
Input Arguments
Bayesian linear regression model storing posterior distribution
characteristics, specified as a conjugateblm or empiricalblm model object returned
by estimate.
When you also specify PriorMdl, then
PosteriorMdl is the posterior distribution composed
of PriorMdl and data. If the
NumPredictors and VarNames
properties of the two models are not equal, plot issues
an error.
Bayesian linear regression model storing prior distribution
characteristics, specified as a conjugateblm, semiconjugateblm, diffuseblm, empiricalblm, customblm, mixconjugateblm, mixsemiconjugateblm, or lassoblm model object
returned by bayeslm.
When you also specify PosteriorMdl, then
PriorMdl is the prior distribution that, when
combined with the data likelihood, forms PosteriorMdl. If
the NumPredictors and VarNames
properties of the two models are not equal, plot issues
an error.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: 'VarNames',["Beta1"; "Beta2"; "Sigma2"] plots the
distributions of regression coefficients corresponding to the names
Beta1 and Beta2 in the
VarNames property of the model object and the disturbance
variance Sigma2.
Parameter names indicating which densities to plot in the figure,
specified as the comma-separated pair consisting of
'VarNames' and a cell vector of character vectors
or string vector. VarNames must include
"Intercept", any name in the
VarNames property of PriorMdl
or PosteriorMdl, or
"Sigma2".
By default, plot chooses
"Intercept" (if an intercept exists in the
model), all regression coefficients, and "Sigma2". If
the model has more than 34 regression coefficients, then
plot chooses the first through the 34th
only.
VarNames is case insensitive.
Tip
If your model contains many variables, then try plotting subsets of the parameters on separate plots for a better view of the distributions.
Example: 'VarNames',["Beta(1)","Beta(2)"]
Data Types: string | cell
Parameter values for density evaluation and plotting, specified as the
comma-separated pair consisting of 'Points' and a
numPoints-dimensional numeric vector or a
numVarNames-dimensional cell vector of numeric
vectors. numPoints is the number of parameters values
that plot evaluates and plots the density.
If
Pointsis a numeric vector, thenplotevaluates and plots the densities of all specified distributions by using its elements (seeVarNames).If
Pointsis a cell vector of numeric vectors, then:numVarNamesmust benumel(, whereVarNames)VarNamesis the value ofVarNames.Cells correspond to the elements of
VarNames.For
j= 1,…,numVarNames,plotevaluates and plots the density of the parameter namedVarNames{j}Points(.j)
By default, plot determines
1000 adequate values at which to evaluate and
plot the density for each parameter.
Example: 'Points',{1:0.1:10 10:0.2:25
1:0.01:2}
Data Types: double | cell
Output Arguments
Parameter values used for density evaluation and plotting, returned as a cell vector of numeric vectors.
Suppose Points and
VarNames are the values of
Points and VarNames,
respectively. If Points is a numeric vector, then
PointsUsed is
repmat({.
Otherwise, Points},numel(VarNames))PointsUsed equals Points.
Cells correspond to the names in VarNames.
Evaluated and plotted posterior densities, returned as a
numVarNames-by-1 cell vector of numeric row vectors.
numVarNames is
numel(, where
VarNames)VarNames is the value of
VarNames. Cells correspond to the names in
VarNames. posteriorDensity
has the same dimensions as priorDensity.
Evaluated and plotted prior densities, returned as a
numVarNames-by-1 cell vector of numeric row vectors.
priorDensity has the same dimensions as
posteriorDensity.
Figure window containing distributions, returned as a figure object.
plot overwrites the figure window that it
produces.
If you rename FigureHandle for a new figure
window, or call figure before calling
plot, then plot continues to
overwrite the current figure. To plot distributions to a different figure
window, change the figure identifier of the current figure window by
renaming its Tag property. For example, to rename the
current figure window called FigureHandle to
newFigure, at the command line,
enter:
FigureHandle.Tag = newFigure;
Limitations
Because improper distributions (distributions with densities
that do not integrate to 1) are not well defined, plot
cannot plot them very well.
More About
A Bayesian linear regression model treats the parameters β and σ2 in the multiple linear regression (MLR) model yt = xtβ + εt as random variables.
For times t = 1,...,T:
yt is the observed response.
xt is a 1-by-(p + 1) row vector of observed values of p predictors. To accommodate a model intercept, x1t = 1 for all t.
β is a (p + 1)-by-1 column vector of regression coefficients corresponding to the variables that compose the columns of xt.
εt is the random disturbance with a mean of zero and Cov(ε) = σ2IT×T, while ε is a T-by-1 vector containing all disturbances. These assumptions imply that the data likelihood is
ϕ(yt;xtβ,σ2) is the Gaussian probability density with mean xtβ and variance σ2 evaluated at yt;.
Before considering the data, you impose a joint prior distribution assumption on (β,σ2). In a Bayesian analysis, you update the distribution of the parameters by using information about the parameters obtained from the likelihood of the data. The result is the joint posterior distribution of (β,σ2) or the conditional posterior distributions of the parameters.
Version History
Introduced in R2017a
See Also
Objects
conjugateblm|semiconjugateblm|diffuseblm|empiricalblm|customblm|mixconjugateblm|mixsemiconjugateblm|lassoblm
Functions
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleziona un sito web
Seleziona un sito web per visualizzare contenuto tradotto dove disponibile e vedere eventi e offerte locali. In base alla tua area geografica, ti consigliamo di selezionare: .
Puoi anche selezionare un sito web dal seguente elenco:
Come ottenere le migliori prestazioni del sito
Per ottenere le migliori prestazioni del sito, seleziona il sito cinese (in cinese o in inglese). I siti MathWorks per gli altri paesi non sono ottimizzati per essere visitati dalla tua area geografica.
Americhe
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)