transform
Syntax
Description
transformedData = transform(remover,Tbl)Tbl according to the transformation in
the disparateImpactRemover object (remover). The
predictor variables and sensitive attribute in Tbl must have the same
names as the variables used to create remover. To see the variable
names, use remover.PredictorNames and
remover.SensitiveAttribute.
To see the fraction of the data transformation used to return
transformedData, use
remover.RepairFraction.
transformedData = transform(remover,X,attribute)X, transformed with respect to the sensitive attribute
attribute.
transformedData = transform(___,Name=Value)RepairFraction name-value argument. A
value of 1 indicates a full transformation, and a value of 0 indicates no
transformation.
Examples
Train a binary classifier, classify test data using the model, and compute the disparate impact for each group in the sensitive attribute. To reduce the disparate impact values, use disparateImpactRemover, and then retrain the binary classifier. Transform the test data set, reclassify the observations, and compute the disparate impact values.
Load the sample data census1994, which contains the training data adultdata and the test data adulttest. The data sets consist of demographic information from the US Census Bureau that can be used to predict whether an individual makes over $50,000 per year. Preview the first few rows of the training data set.
load census1994
head(adultdata) age workClass fnlwgt education education_num marital_status occupation relationship race sex capital_gain capital_loss hours_per_week native_country salary
___ ________________ __________ _________ _____________ _____________________ _________________ _____________ _____ ______ ____________ ____________ ______________ ______________ ______
39 State-gov 77516 Bachelors 13 Never-married Adm-clerical Not-in-family White Male 2174 0 40 United-States <=50K
50 Self-emp-not-inc 83311 Bachelors 13 Married-civ-spouse Exec-managerial Husband White Male 0 0 13 United-States <=50K
38 Private 2.1565e+05 HS-grad 9 Divorced Handlers-cleaners Not-in-family White Male 0 0 40 United-States <=50K
53 Private 2.3472e+05 11th 7 Married-civ-spouse Handlers-cleaners Husband Black Male 0 0 40 United-States <=50K
28 Private 3.3841e+05 Bachelors 13 Married-civ-spouse Prof-specialty Wife Black Female 0 0 40 Cuba <=50K
37 Private 2.8458e+05 Masters 14 Married-civ-spouse Exec-managerial Wife White Female 0 0 40 United-States <=50K
49 Private 1.6019e+05 9th 5 Married-spouse-absent Other-service Not-in-family Black Female 0 0 16 Jamaica <=50K
52 Self-emp-not-inc 2.0964e+05 HS-grad 9 Married-civ-spouse Exec-managerial Husband White Male 0 0 45 United-States >50K
Each row contains the demographic information for one adult. The last column salary shows whether a person has a salary less than or equal to $50,000 per year or greater than $50,000 per year.
Remove observations from adultdata and adulttest that contain missing values.
adultdata = rmmissing(adultdata); adulttest = rmmissing(adulttest);
Specify the continuous numeric predictors to use for model training.
predictors = ["age","education_num","capital_gain","capital_loss", ... "hours_per_week"];
Train an ensemble classifier using the training set adultdata. Specify salary as the response variable and fnlwgt as the observation weights. Because the training set is imbalanced, use the RUSBoost algorithm. After training the model, predict the salary (class label) of the observations in the test set adulttest.
rng("default") % For reproducibility mdl = fitcensemble(adultdata,"salary",Weights="fnlwgt", ... PredictorNames=predictors,Method="RUSBoost"); labels = predict(mdl,adulttest);
Transform the training set predictors by using the race sensitive attribute.
[remover,newadultdata] = disparateImpactRemover(adultdata, ... "race",PredictorNames=predictors); remover
remover =
disparateImpactRemover with properties:
RepairFraction: 1
PredictorNames: {'age' 'education_num' 'capital_gain' 'capital_loss' 'hours_per_week'}
SensitiveAttribute: 'race'
remover is a disparateImpactRemover object, which contains the transformation of the remover.PredictorNames predictors with respect to the remover.SensitiveAttribute variable.
Apply the same transformation stored in remover to the test set predictors. Note: You must transform both the training and test data sets before passing them to a classifier.
newadulttest = transform(remover,adulttest, ...
PredictorNames=predictors);Train the same type of ensemble classifier as mdl, but use the transformed predictor data. As before, predict the salary (class label) of the observations in the test set adulttest.
rng("default") % For reproducibility newMdl = fitcensemble(newadultdata,"salary",Weights="fnlwgt", ... PredictorNames=predictors,Method="RUSBoost"); newLabels = predict(newMdl,newadulttest);
Compare the disparate impact values for the predictions made by the original model (mdl) and the predictions made by the model trained with the transformed data (newMdl). For each group in the sensitive attribute, the disparate impact value is the proportion of predictions in that group with a positive class value () divided by the proportion of predictions in the reference group with a positive class value (). An ideal classifier makes predictions where, for each group, is close to (that is, where the disparate impact value is close to 1).
Compute the disparate impact values for the mdl predictions and the newMdl predictions by using fairnessMetrics. Include the observation weights. You can use the report object function to display bias metrics, such as disparate impact, that are stored in the metricsResults object.
metricsResults = fairnessMetrics(adulttest,"salary", ... SensitiveAttributeNames="race",Predictions=[labels,newLabels], ... Weights="fnlwgt",ModelNames=["Original Model","New Model"]); metricsResults.PositiveClass
ans = categorical
>50K
metricsResults.ReferenceGroup
ans = 'White'
report(metricsResults,BiasMetrics="DisparateImpact")ans=5×5 table
Metrics SensitiveAttributeNames Groups Original Model New Model
_______________ _______________________ __________________ ______________ _________
DisparateImpact race Amer-Indian-Eskimo 0.41702 0.92804
DisparateImpact race Asian-Pac-Islander 1.719 0.9697
DisparateImpact race Black 0.60571 0.66629
DisparateImpact race Other 0.66958 0.86039
DisparateImpact race White 1 1
For the mdl predictions, several of the disparate impact values are below the industry standard of 0.8, and one value is above 1.25. These values indicate bias in the predictions with respect to the positive class >50K and the sensitive attribute race.
The disparate impact values for the newMdl predictions are closer to 1 than the disparate impact values for the mdl predictions. One value is still below 0.8.
Visually compare the disparate impact values by using the bar graph returned by the plot object function.
plot(metricsResults,"DisparateImpact")
The disparateImpactRemover function seems to have improved the model predictions on the test set with respect to the disparate impact metric.
Check whether the transformed predictors negatively affect the accuracy of the model predictions. Compute the accuracy of the test set predictions for the two models mdl and newMdl.
accuracy = 1-loss(mdl,adulttest,"salary")accuracy = 0.8024
newAccuracy = 1-loss(newMdl,newadulttest,"salary")newAccuracy = 0.7955
The model trained using the transformed predictors (newMdl) achieves similar test set accuracy compared to the model trained with the original predictors (mdl).
Specify the extent of the transformation of the continuous numeric predictors with respect to a sensitive attribute. Use the RepairFraction name-value argument of the disparateImpactRemover function.
Load the patients data set, which contains medical information for 100 patients. Convert the Gender and Smoker variables to categorical variables. Specify the descriptive category names Smoker and Nonsmoker rather than 1 and 0.
load patients Gender = categorical(Gender); Smoker = categorical(Smoker,logical([1 0]), ... ["Smoker","Nonsmoker"]);
Create a matrix containing the continuous predictors Diastolic and Systolic.
X = [Diastolic,Systolic];
Find the observations in the two groups of the sensitive attribute Gender.
femaleIdx = Gender=="Female"; maleIdx = Gender=="Male"; femaleX = X(femaleIdx,:); maleX = X(maleIdx,:);
Transform the Diastolic and Systolic predictors in X by using the Gender sensitive attribute. Specify a repair fraction of 0.5. Note that a value of 1 indicates a full transformation, and a value of 0 indicates no transformation.
[remover,newX50] = disparateImpactRemover(X,Gender, ...
RepairFraction=0.5);
femaleNewX50 = newX50(femaleIdx,:);
maleNewX50 = newX50(maleIdx,:);Fully transform the predictor variables by using the transform object function of the remover object.
newX100 = transform(remover,X,Gender,RepairFraction=1); femaleNewX100 = newX100(femaleIdx,:); maleNewX100 = newX100(maleIdx,:);
Visualize the difference in the Diastolic distributions between the original values in X, the partially repaired values in newX50, and the fully transformed values in newX100. Compute and display the probability density estimates by using the ksdensity function.
t = tiledlayout(1,3); title(t,"Diastolic Distributions with Different " + ... "Repair Fractions") xlabel(t,"Diastolic") ylabel(t,"Density Estimate") nexttile ksdensity(femaleX(:,1)) hold on ksdensity(maleX(:,1)) hold off title("Fraction=0") ylim([0,0.07]) nexttile ksdensity(femaleNewX50{:,1}) hold on ksdensity(maleNewX50{:,1}) hold off title("Fraction=0.5") ylim([0,0.07]) nexttile ksdensity(femaleNewX100{:,1}) hold on ksdensity(maleNewX100{:,1}) hold off title("Fraction=1") ylim([0,0.07]) legend(["Female","Male"],Location="eastoutside")
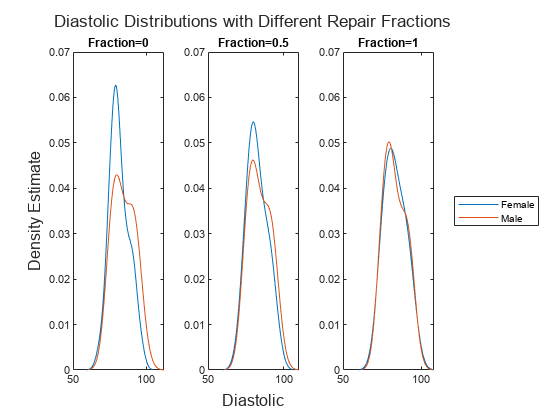
As the repair fraction increases, the disparateImpactRemover function transforms the values in the Diastolic predictor variable so that the distribution of Female values and the distribution of Male values become more similar.
Input Arguments
Predictor data transformer, specified as a disparateImpactRemover object. For a new data set, the
transform object function transforms the
remover.PredictorNames predictor variables with respect to the
sensitive attribute specified by remover.SensitiveAttribute.
Note that if remover.SensitiveAttribute is a variable rather than
the name of a variable, then transform does not use the stored
sensitive attribute values when transforming new data. The function uses the values in
attribute instead.
Data set, specified as a table. Each row of Tbl corresponds to
one observation, and each column corresponds to one variable. If you use a table when
creating the disparateImpactRemover object, then you must use a table
when using the transform object function. The table must include
all required predictor variables and the sensitive attribute. The table can include
additional variables, such as the response variable. Multicolumn variables and cell
arrays other than cell arrays of character vectors are not allowed.
Data Types: table
Predictor data, specified as a numeric matrix. Each row of X
corresponds to one observation, and each column corresponds to one predictor variable.
If you use a matrix when creating the disparateImpactRemover object,
then you must use a matrix when using the transform object
function. X and attribute must have the same
number of rows.
Data Types: single | double
Sensitive attribute, specified as a numeric column vector, logical column vector, character array, string array, cell array of character vectors, or categorical column vector.
The data type of
attributemust be the same as the data type ofremover.SensitiveAttribute. (The software treats string arrays as cell arrays of character vectors.)The distinct classes in
attributemust be a subset of the classes inremover.SensitiveAttribute.If
attributeis an array, then each row of the array must correspond to a group in the sensitive attribute.attributeandXmust have the same number of rows.
Data Types: single | double | logical | char | string | cell | categorical
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: transform(remover,Tbl,RepairFraction=1,PredictorNames=["Diastolic","Systolic"])
specifies to transform fully the Diastolic and
Systolic variables in the table Tbl by using the
transformation stored in remover.
Names of the predictor variables to transform, specified as a string array of
unique names or cell array of unique character vectors. The predictor variable names
must be a subset of the names stored in
remover.PredictorNames.
Example: PredictorNames=["SepalLength","SepalWidth","PetalLength","PetalWidth"]
Data Types: string | cell
Fraction of the data transformation, specified as a numeric scalar in the range [0,1]. A value of 1 indicates a full transformation, and a value of 0 indicates no transformation.
A greater repair fraction can result in a greater loss in model prediction accuracy.
Example: RepairFraction=0.75
Data Types: single | double
Output Arguments
Transformed predictor data, returned as a table or numeric matrix. Note that
transformedData can include the sensitive attribute. After you
use the disparateImpactRemover function, avoid using the sensitive
attribute as a separate predictor when training your model.
For more information on how disparateImpactRemover transforms
predictor data, see Algorithms.
Version History
Introduced in R2022b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleziona un sito web
Seleziona un sito web per visualizzare contenuto tradotto dove disponibile e vedere eventi e offerte locali. In base alla tua area geografica, ti consigliamo di selezionare: .
Puoi anche selezionare un sito web dal seguente elenco:
Come ottenere le migliori prestazioni del sito
Per ottenere le migliori prestazioni del sito, seleziona il sito cinese (in cinese o in inglese). I siti MathWorks per gli altri paesi non sono ottimizzati per essere visitati dalla tua area geografica.
Americhe
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)